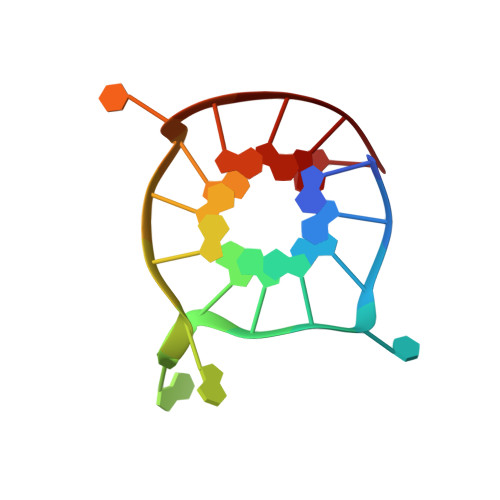
Overlapping but distinct: a new model for G-quadruplex biochemical specificity.
Volek, M., Kolesnikova, S., Svehlova, K., Srb, P., Sgallova, R., Streckerova, T., Redondo, J.A., Veverka, V., Curtis, E.A.(2021) Nucleic Acids Res 49: 1816-1827
- PubMed: 33544841 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkab037
- Primary Citation Related Structures:
6YY4 - PubMed Abstract:
G-quadruplexes are noncanonical nucleic acid structures formed by stacked guanine tetrads. They are capable of a range of functions and thought to play widespread biological roles. This diversity raises an important question: what determines the biochemical specificity of G-quadruplex structures? The answer is particularly important from the perspective of biological regulation because genomes can contain hundreds of thousands of G-quadruplexes with a range of functions. Here we analyze the specificity of each sequence in a 496-member library of variants of a reference G-quadruplex with respect to five functions. Our analysis shows that the sequence requirements of G-quadruplexes with these functions are different from one another, with some mutations altering biochemical specificity by orders of magnitude. Mutations in tetrads have larger effects than mutations in loops, and changes in specificity are correlated with changes in multimeric state. To complement our biochemical data we determined the solution structure of a monomeric G-quadruplex from the library. The stacked and accessible tetrads rationalize why monomers tend to promote a model peroxidase reaction and generate fluorescence. Our experiments support a model in which the sequence requirements of G-quadruplexes with different functions are overlapping but distinct. This has implications for biological regulation, bioinformatics, and drug design.
- Institute of Organic Chemistry and Biochemistry of the Czech Academy of Sciences, Prague 166 10, Czech Republic.
Organizational Affiliation: