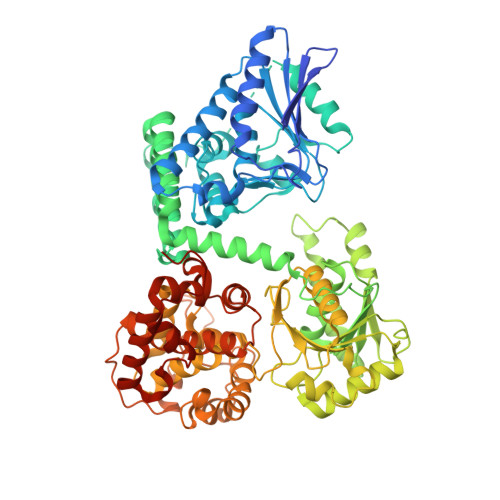
Structural basis for different membrane-binding properties of E. coli anaerobic and human mitochondrial beta-oxidation trifunctional enzymes
Sah-Teli, S.K., Pinkas, M., Hynonen, M.J., Butcher, S.J., Wierenga, R.K., Novacek, J., Venkatesan, R.(2023) Structure