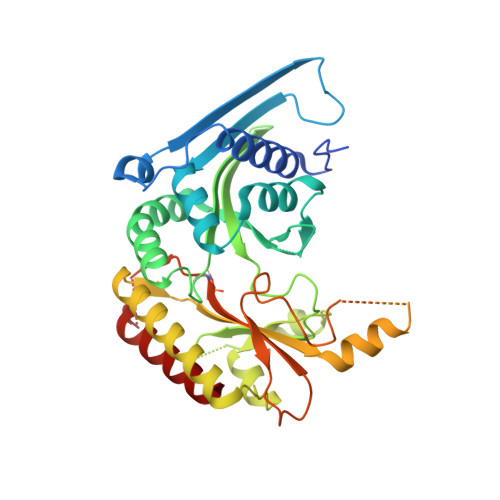
Discovery and Characterization of the Potent and Highly Selective 1,7-Naphthyridine-Based Inhibitors BAY-091 and BAY-297 of the Kinase PIP4K2A.
Wortmann, L., Brauer, N., Holton, S.J., Irlbacher, H., Weiske, J., Lechner, C., Meier, R., Karen, J., Sioberg, C.B., Putter, V., Christ, C.D., Ter Laak, A., Lienau, P., Lesche, R., Nicke, B., Cheung, S.H., Bauser, M., Haegebarth, A., von Nussbaum, F., Mumberg, D., Lemos, C.(2021) J Med Chem 64: 15883-15911
- PubMed: 34699202 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.1c01245
- Primary Citation Related Structures:
6YM3, 6YM4, 6YM5 - PubMed Abstract:
PIP4K2A is an insufficiently studied type II lipid kinase that catalyzes the conversion of phosphatidylinositol-5-phosphate (PI5P) into phosphatidylinositol 4,5-bisphosphate (PI4,5P 2 ). The involvement of PIP4K2A/B in cancer has been suggested, particularly in the context of p53 mutant/null tumors. PIP4K2A/B depletion has been shown to induce tumor growth inhibition, possibly due to hyperactivation of AKT and reactive oxygen species-mediated apoptosis. Herein, we report the identification of the novel potent and highly selective inhibitors BAY-091 and BAY-297 of the kinase PIP4K2A by high-throughput screening and subsequent structure-based optimization. Cellular target engagement of BAY-091 and BAY-297 was demonstrated using cellular thermal shift assay technology. However, inhibition of PIP4K2A with BAY-091 or BAY-297 did not translate into the hypothesized mode of action and antiproliferative activity in p53-deficient tumor cells. Therefore, BAY-091 and BAY-297 serve as valuable chemical probes to study PIP4K2A signaling and its involvement in pathophysiological conditions such as cancer.
- Bayer AG, Research & Development, Pharmaceuticals, 13353 Berlin, Germany.
Organizational Affiliation: