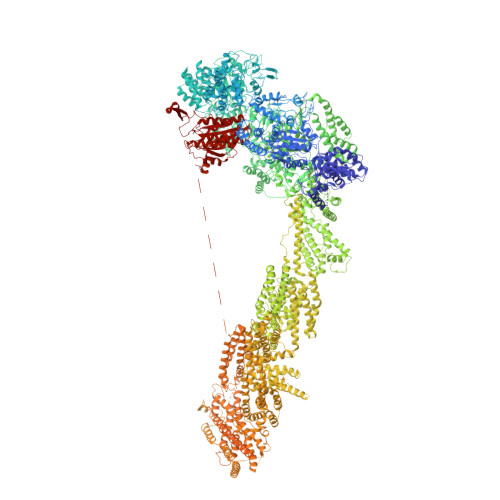
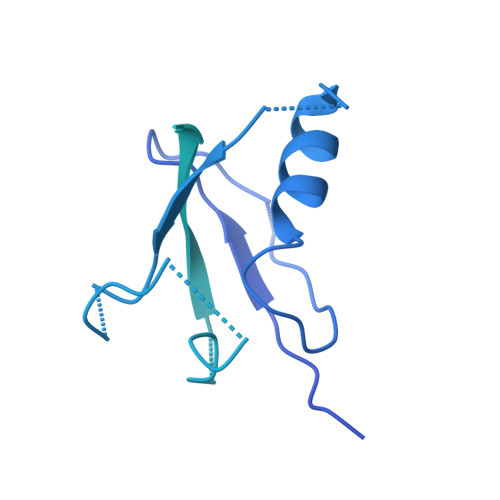
Construction of the Central Protuberance and L1 Stalk during 60S Subunit Biogenesis.
Kater, L., Mitterer, V., Thoms, M., Cheng, J., Berninghausen, O., Beckmann, R., Hurt, E.(2020) Mol Cell 79: 615-628.e5
- PubMed: 32668200 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2020.06.032
- Primary Citation Related Structures:
6YLE, 6YLF, 6YLG, 6YLH, 6YLX, 6YLY - PubMed Abstract:
Ribosome assembly is driven by numerous assembly factors, including the Rix1 complex and the AAA ATPase Rea1. These two assembly factors catalyze 60S maturation at two distinct states, triggering poorly understood large-scale structural transitions that we analyzed by cryo-electron microscopy. Two nuclear pre-60S intermediates were discovered that represent previously unknown states after Rea1-mediated removal of the Ytm1-Erb1 complex and reveal how the L1 stalk develops from a pre-mature nucleolar to a mature-like nucleoplasmic state. A later pre-60S intermediate shows how the central protuberance arises, assisted by the nearby Rix1-Rea1 machinery, which was solved in its pre-ribosomal context to molecular resolution. This revealed a Rix1 2 -Ipi3 2 tetramer anchored to the pre-60S via Ipi1, strategically positioned to monitor this decisive remodeling. These results are consistent with a general underlying principle that temporarily stabilized immature RNA domains are successively remodeled by assembly factors, thereby ensuring failsafe assembly progression.
- Gene Center Munich and Center of Integrated Protein Science-Munich (CiPS-M), Department of Biochemistry, Feodor-Lynen-Str. 25, University of Munich, 81377 Munich, Germany.
Organizational Affiliation: