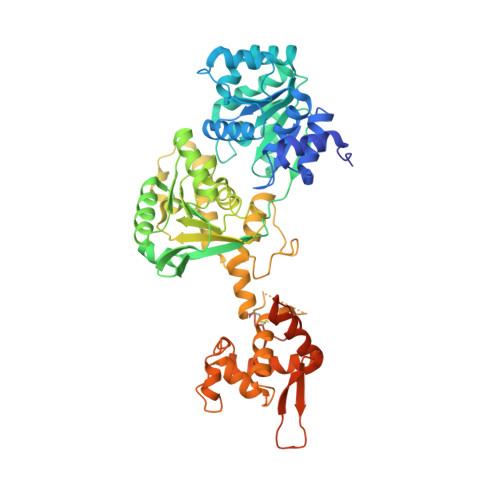
Structure of the helicase core of Werner helicase, a key target in microsatellite instability cancers.
Newman, J.A., Gavard, A.E., Lieb, S., Ravichandran, M.C., Hauer, K., Werni, P., Geist, L., Bottcher, J., Engen, J.R., Rumpel, K., Samwer, M., Petronczki, M., Gileadi, O.(2021) Life Sci Alliance 4
- PubMed: 33199508 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.26508/lsa.202000795
- Primary Citation Related Structures:
6YHR - PubMed Abstract:
Loss of WRN, a DNA repair helicase, was identified as a strong vulnerability of microsatellite instable (MSI) cancers, making WRN a promising drug target. We show that ATP binding and hydrolysis are required for genome integrity and viability of MSI cancer cells. We report a 2.2-Å crystal structure of the WRN helicase core (517-1,093), comprising the two helicase subdomains and winged helix domain but not the HRDC domain or nuclease domains. The structure highlights unusual features. First, an atypical mode of nucleotide binding that results in unusual relative positioning of the two helicase subdomains. Second, an additional β-hairpin in the second helicase subdomain and an unusual helical hairpin in the Zn 2+ binding domain. Modelling of the WRN helicase in complex with DNA suggests roles for these features in the binding of alternative DNA structures. NMR analysis shows a weak interaction between the HRDC domain and the helicase core, indicating a possible biological role for this association. Together, this study will facilitate the structure-based development of inhibitors against WRN helicase.
- Structural Genomics Consortium, University of Oxford, Oxford, UK.
Organizational Affiliation: