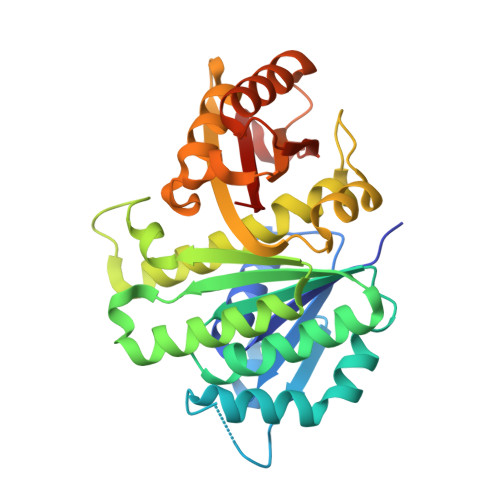
Conformational Flexibility of A Highly Conserved Helix Controls Cryptic Pocket Formation in FtsZ.
Alnami, A., Norton, R.S., Pena, H.P., Haider, S., Kozielski, F.(2021) J Mol Biology 433: 167061-167061
- PubMed: 34023403 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2021.167061
- Primary Citation Related Structures:
6Y1U, 6Y1V, 6YM1, 6YM9 - PubMed Abstract:
Mycobacterium tuberculosis is responsible for more than 1.6 million deaths each year. One potential antibacterial target in M. tuberculosis is filamentous temperature sensitive protein Z (FtsZ), which is the bacterial homologue of mammalian tubulin, a validated cancer target. M. tuberculosis FtsZ function is essential, with its inhibition leading to arrest of cell division, elongation of the bacterial cell and eventual cell death. However, the development of potent inhibitors against FtsZ has been a challenge owing to the lack of structural information. Here we report multiple crystal structures of M. tuberculosis FtsZ in complex with a coumarin analogue. The 4-hydroxycoumarin binds exclusively to two novel cryptic pockets in nucleotide-free FtsZ, but not to the binary FtsZ-GTP or GDP complexes. Our findings provide a detailed understanding of the molecular basis for cryptic pocket formation, controlled by the conformational flexibility of the H7 helix, and thus reveal an important structural and mechanistic rationale for coumarin antibacterial activity.
- Department of Pharmaceutical and Biological Chemistry, School of Pharmacy, University College London, 29-39 Brunswick Square, London WC1N 1AX, UK.
Organizational Affiliation: