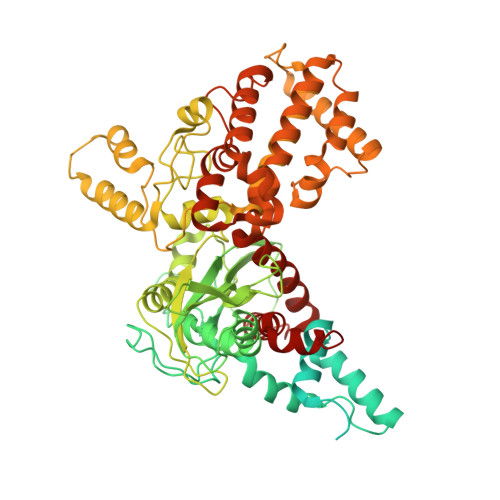
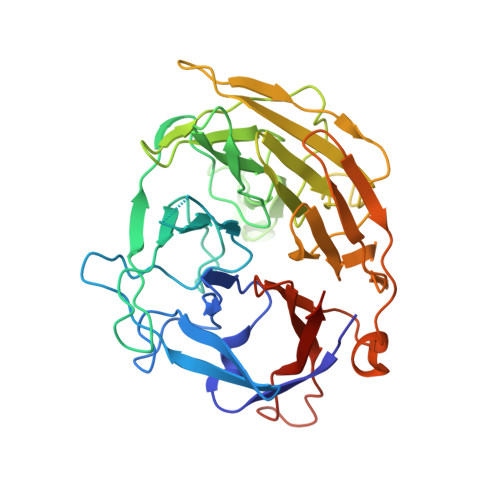
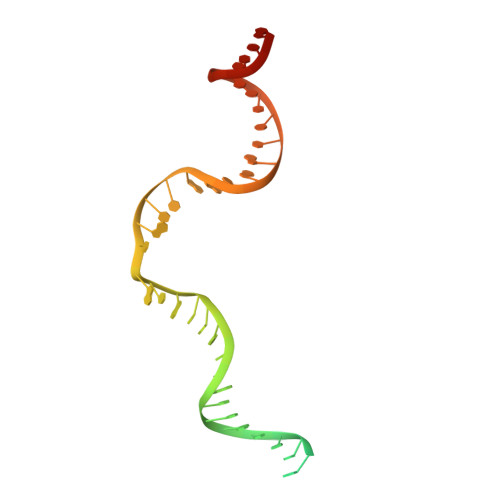
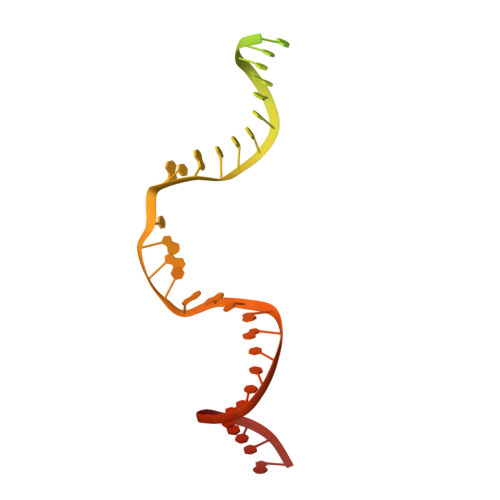Structural basis for the activation and suppression of transposition during evolution of the RAG recombinase.
Zhang, Y., Corbett, E., Wu, S., Schatz, D.G.(2020) EMBO J 39: e105857-e105857
- PubMed: 32945578 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.15252/embj.2020105857
- Primary Citation Related Structures:
6XNX, 6XNY, 6XNZ - PubMed Abstract:
Jawed vertebrate adaptive immunity relies on the RAG1/RAG2 (RAG) recombinase, a domesticated transposase, for assembly of antigen receptor genes. Using an integration-activated form of RAG1 with methionine at residue 848 and cryo-electron microscopy, we determined structures that capture RAG engaged with transposon ends and U-shaped target DNA prior to integration (the target capture complex) and two forms of the RAG strand transfer complex that differ based on whether target site DNA is annealed or dynamic. Target site DNA base unstacking, flipping, and melting by RAG1 methionine 848 explain how this residue activates transposition, how RAG can stabilize sharp bends in target DNA, and why replacement of residue 848 by arginine during RAG domestication led to suppression of transposition activity. RAG2 extends a jawed vertebrate-specific loop to interact with target site DNA, and functional assays demonstrate that this loop represents another evolutionary adaptation acquired during RAG domestication to inhibit transposition. Our findings identify mechanistic principles of the final step in cut-and-paste transposition and the molecular and structural logic underlying the transformation of RAG from transposase to recombinase.
- Department of Immunobiology, Yale School of Medicine, New Haven, CT, USA.
Organizational Affiliation: