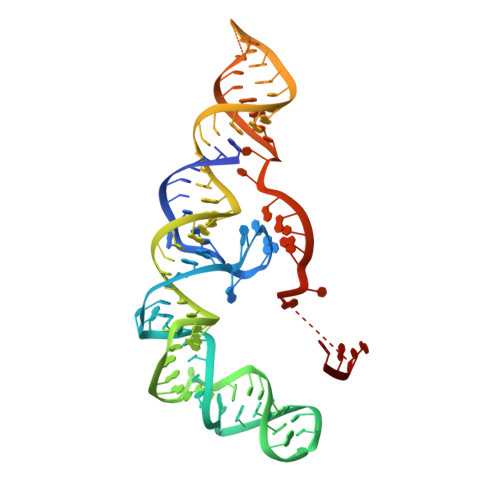
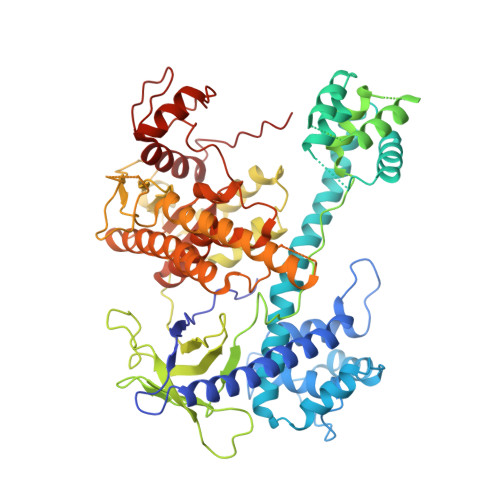
Cryo-EM structure of the RNA-guided ribonuclease Cas12g.
Li, Z., Zhang, H., Xiao, R., Han, R., Chang, L.(2021) Nat Chem Biol 17: 387-393
- PubMed: 33495647 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41589-020-00721-2
- Primary Citation Related Structures:
6XMF, 6XMG - PubMed Abstract:
Cas12g, the type V-G CRISPR-Cas effector, is an RNA-guided ribonuclease that targets single-stranded RNA substrate. The CRISPR-Cas12g system offers a potential platform for transcriptome engineering and diagnostic applications. We determined the structures of Cas12g-guide RNA complexes in the absence and presence of target RNA by cryo-EM to a resolution of 3.1 Å and 4.8 Å, respectively. Cas12g adopts a bilobed structure with miniature REC2 and Nuc domains, whereas the guide RNAs fold into a flipped 'F' shape, which is primarily recognized by the REC lobe. Target RNA and the CRISPR RNA (crRNA) guide form a duplex that inserts into the central cavity between the REC and NUC lobes, inducing conformational changes in both lobes to activate Cas12g. The structural insights would facilitate the development of Cas12g-based applications.
- Department of Biological Sciences, Purdue University, West Lafayette, IN, USA.
Organizational Affiliation: