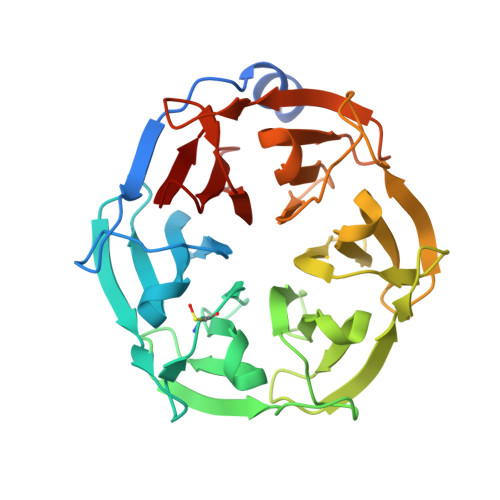
Costameric integrin and sarcoglycan protein levels are altered in a Drosophila model for Limb-girdle muscular dystrophy type 2H.
Bawa, S., Gameros, S., Baumann, K., Brooks, D.S., Kollhoff, J.A., Zolkiewski, M., Re Cecconi, A.D., Panini, N., Russo, M., Piccirillo, R., Johnson, D.K., Kashipathy, M.M., Battaile, K.P., Lovell, S., Bouyain, S.E.A., Kawakami, J., Geisbrecht, E.R.(2021) Mol Biol Cell 32: 260-273
- PubMed: 33296226 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1091/mbc.E20-07-0453
- Primary Citation Related Structures:
6XG7 - PubMed Abstract:
Mutations in two different domains of the ubiquitously expressed TRIM32 protein give rise to two clinically separate diseases, one of which is Limb-girdle muscular dystrophy type 2H (LGMD2H). Uncovering the muscle-specific role of TRIM32 in LGMD2H pathogenesis has proven difficult, as neurogenic phenotypes, independent of LGMD2H pathology, are present in TRIM32 KO mice. We previously established a platform to study LGMD2H pathogenesis using Drosophila melanogaster as a model. Here we show that LGMD2H disease-causing mutations in the NHL domain are molecularly and structurally conserved between fly and human TRIM32. Furthermore, transgenic expression of a subset of myopathic alleles (R394H, D487N, and 520fs) induce myofibril abnormalities, altered nuclear morphology, and reduced TRIM32 protein levels, mimicking phenotypes in patients afflicted with LGMD2H. Intriguingly, we also report for the first time that the protein levels of βPS integrin and sarcoglycan δ, both core components of costameres, are elevated in TRIM32 disease-causing alleles. Similarly, murine myoblasts overexpressing a catalytically inactive TRIM32 mutant aberrantly accumulate α- and β-dystroglycan and α-sarcoglycan. We speculate that the stoichiometric loss of costamere components disrupts costamere complexes to promote muscle degeneration.
- Department of Biochemistry and Molecular Biophysics, Kansas State University, Manhattan, KS 66506.
Organizational Affiliation: