An orthogonal seryl-tRNA synthetase/tRNA pair for noncanonical amino acid mutagenesis in Escherichia coli.
Zambaldo, C., Koh, M., Nasertorabi, F., Han, G.W., Chatterjee, A., Stevens, R.C., Schultz, P.G.(2020) Bioorg Med Chem 28: 115662-115662
- PubMed: 33069069 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.bmc.2020.115662
- Primary Citation Related Structures:
6X94 - PubMed Abstract:
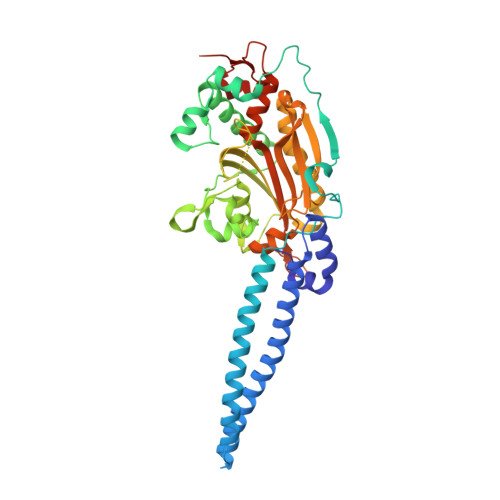
We report the development of the orthogonal amber-suppressor pair Archaeoglobus fulgidus seryl-tRNA (Af-tRNA Ser )/Methanosarcina mazei seryl-tRNA synthetase (MmSerRS) in Escherichia coli. Furthermore, the crystal structure of MmSerRS was solved at 1.45 Å resolution, which should enable structure-guided engineering of its active site to genetically encode small, polar noncanonical amino acids (ncAAs).
- Department of Chemistry and Skaggs Institute for Chemical Biology, The Scripps Research Institute, 10550 N. Torrey Pines Road, La Jolla, CA 92037, United States.
Organizational Affiliation: