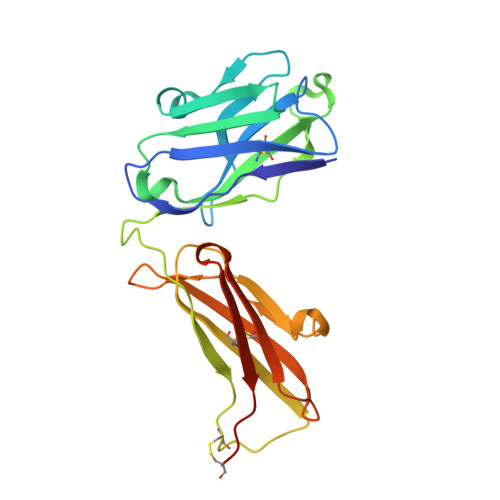
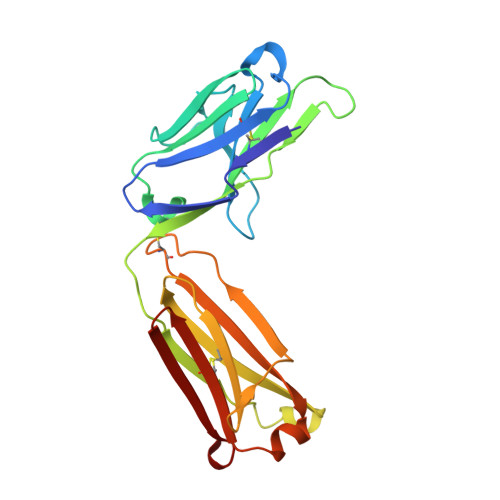
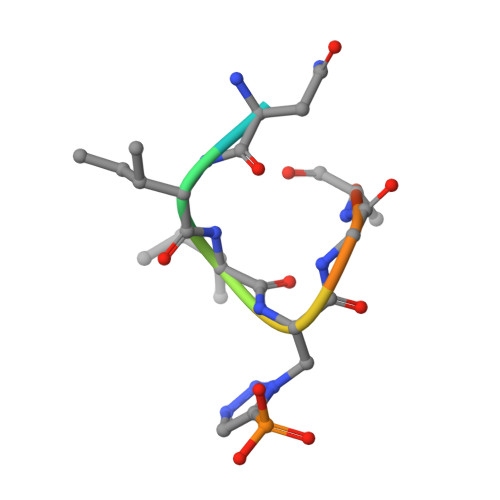
Structural basis for differential recognition of phosphohistidine-containing peptides by 1-pHis and 3-pHis monoclonal antibodies.
Kalagiri, R., Stanfield, R.L., Meisenhelder, J., La Clair, J.J., Fuhs, S.R., Wilson, I.A., Hunter, T.(2021) Proc Natl Acad Sci U S A 118
- PubMed: 33547238 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2010644118
- Primary Citation Related Structures:
6X1S, 6X1T, 6X1U, 6X1V, 6X1W - PubMed Abstract:
In 2015, monoclonal antibodies (mAbs) that selectively recognize the 1-pHis or 3-pHis isoforms of phosphohistidine were developed by immunizing rabbits with degenerate Ala/Gly peptides containing the nonhydrolyzable phosphohistidine (pHis) analog- phosphotriazolylalanine (pTza). Here, we report structures of five rabbit mAbs bound to cognate pTza peptides: SC1-1 and SC50-3 that recognize 1-pHis, and their 3-pHis-specific counterparts, SC39-4, SC44-8, and SC56-2. These cocrystal structures provide insights into the binding modes of the pTza phosphate group that are distinct for the 1- and 3-pHis mAbs with the selectivity arising from specific contacts with the phosphate group and triazolyl ring. The mode of phosphate recognition in the 3-pHis mAbs recapitulates the Walker A motif, as present in kinases. The complementarity-determining regions (CDRs) of four of the Fabs interact with the peptide backbone rather than peptide side chains, thus conferring sequence independence, whereas SC44-8 shows a proclivity for binding a GpHAGA motif mediated by a sterically complementary CDRL3 loop. Specific hydrogen bonding with the triazolyl ring precludes recognition of pTyr and other phosphoamino acids by these mAbs. Kinetic binding experiments reveal that the affinity of pHis mAbs for pHis and pTza peptides is submicromolar. Bound pHis mAbs also shield the pHis peptides from rapid dephosphorylation. The epitope-paratope interactions illustrate how these anti-pHis antibodies are useful for a wide range of research techniques and this structural information can be utilized to improve the specificity and affinity of these antibodies toward a variety of pHis substrates to understand the role of histidine phosphorylation in healthy and diseased states.
- Molecular and Cell Biology Laboratory, Salk Institute for Biological Studies, La Jolla, CA 92037.
Organizational Affiliation: