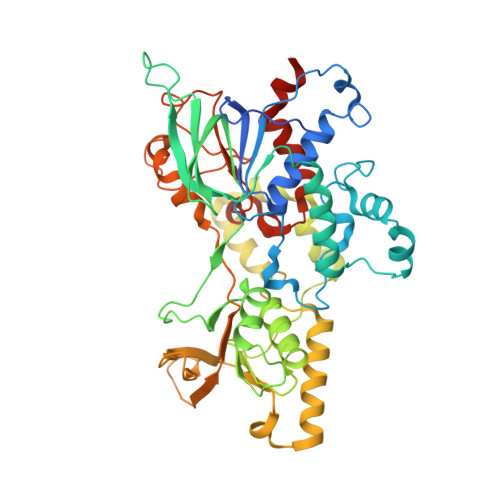
Trapping conformational states of a flavin-dependent N -monooxygenase in crystallo reveals protein and flavin dynamics.
Campbell, A.C., Stiers, K.M., Martin Del Campo, J.S., Mehra-Chaudhary, R., Sobrado, P., Tanner, J.J.(2020) J Biological Chem 295: 13239-13249
- PubMed: 32723870 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA120.014750
- Primary Citation Related Structures:
6X0H, 6X0I, 6X0J, 6X0K - PubMed Abstract:
The siderophore biosynthetic enzyme A (SidA) ornithine hydroxylase from Aspergillus fumigatus is a fungal disease drug target involved in the production of hydroxamate-containing siderophores, which are used by the pathogen to sequester iron. SidA is an N -monooxygenase that catalyzes the NADPH-dependent hydroxylation of l-ornithine through a multistep oxidative mechanism, utilizing a C4a-hydroperoxyflavin intermediate. Here we present four new crystal structures of SidA in various redox and ligation states, including the first structure of oxidized SidA without NADP(H) or l-ornithine bound (resting state). The resting state structure reveals a new out active site conformation characterized by large rotations of the FAD isoalloxazine around the C1-'C2' and N10-C1' bonds, coupled to a 10-Å movement of the Tyr-loop. Additional structures show that either flavin reduction or the binding of NADP(H) is sufficient to drive the FAD to the in conformation. The structures also reveal protein conformational changes associated with the binding of NADP(H) and l-ornithine. Some of these residues were probed using site-directed mutagenesis. Docking was used to explore the active site of the out conformation. These calculations identified two potential ligand-binding sites. Altogether, our results provide new information about conformational dynamics in flavin-dependent monooxygenases. Understanding the different active site conformations that appear during the catalytic cycle may allow fine-tuning of inhibitor discovery efforts.
- Department of Biochemistry, University of Missouri, Columbia, Missouri, USA.
Organizational Affiliation: