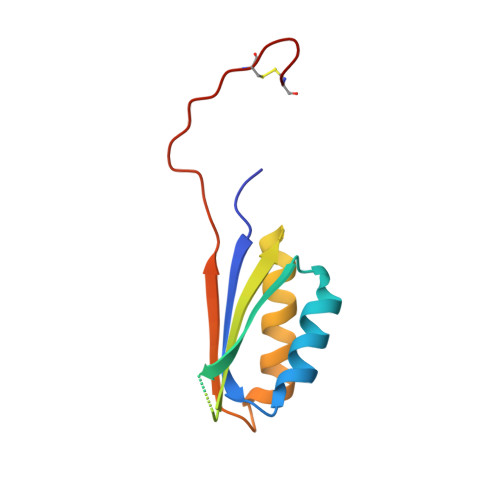
Tetragonal crystal form of the cyanobacterial bicarbonate-transporter regulator SbtB from Synechocystis sp. PCC 6803.
Bu, G., Simmons, C.R., Nielsen, D.R., Nannenga, B.L.(2020) Acta Crystallogr F Struct Biol Commun 76: 438-443
- PubMed: 32880592 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X20010523
- Primary Citation Related Structures:
6WUE - PubMed Abstract:
The P II -like protein SbtB has been identified as a regulator of SbtA, which is one of the key bicarbonate transporters in cyanobacteria. While SbtB from Synechocystis sp. PCC 6803 has previously been shown to be a trimer, a new crystal form is reported here which crystallizes in what is thought to be a non-native tetramer in the crystal, with the C-terminus in an extended conformation. The crystal structure shows the formation of an intermolecular disulfide bond at Cys94 between SbtB monomers, which may stabilize this conformation in the crystal. This motivates the need for future studies to investigate the potential role that the oxidation and reduction of these cysteines may play in the activation and/or function of SbtB.
- Chemical Engineering, School for Engineering of Matter, Transport and Energy, Arizona State University, Tempe, AZ 85287, USA.
Organizational Affiliation: