Structure-Based Drug Design of Phenazopyridine Derivatives as Inhibitors of Rev1 Interactions in Translesion Synthesis.
McPherson, K.S., Zaino, A.M., Dash, R.C., Rizzo, A.A., Li, Y., Hao, B., Bezsonova, I., Hadden, M.K., Korzhnev, D.M.(2021) ChemMedChem 16: 1126-1132
- PubMed: 33314657 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/cmdc.202000893
- Primary Citation Related Structures:
6WS0, 6WS5 - PubMed Abstract:
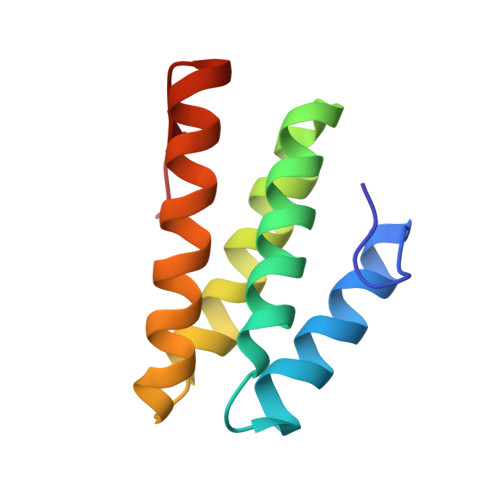
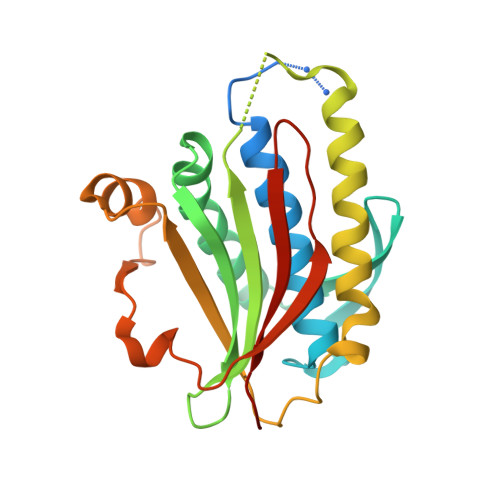
Rev1 is a protein scaffold of the translesion synthesis (TLS) pathway, which employs low-fidelity DNA polymerases for replication of damaged DNA. The TLS pathway helps cancers tolerate DNA damage induced by genotoxic chemotherapy, and increases mutagenesis in tumors, thus accelerating the onset of chemoresistance. TLS inhibitors have emerged as potential adjuvant drugs to enhance the efficacy of first-line chemotherapy, with the majority of reported inhibitors targeting protein-protein interactions (PPIs) of the Rev1 C-terminal domain (Rev1-CT). We previously identified phenazopyridine (PAP) as a scaffold to disrupt Rev1-CT PPIs with Rev1-interacting regions (RIRs) of TLS polymerases. To explore the structure-activity relationships for this scaffold, we developed a protocol for co-crystallization of compounds that target the RIR binding site on Rev1-CT with a triple Rev1-CT/Rev7 R124A /Rev3-RBM1 complex, and solved an X-ray crystal structure of Rev1-CT bound to the most potent PAP analogue. The structure revealed an unexpected binding pose of the compound and informed changes to the scaffold to improve its affinity for Rev1-CT. We synthesized eight additional PAP derivatives, with modifications to the scaffold driven by the structure, and evaluated their binding to Rev1-CT by microscale thermophoresis (MST). Several second-generation PAP derivatives showed an affinity for Rev1-CT that was improved by over an order of magnitude, thereby validating the structure-based assumptions that went into the compound design.
- Department of Molecular Biology and Biophysics, University of Connecticut Health Center, 263 Farmington Ave., Farmington, CT 06030, USA.
Organizational Affiliation: