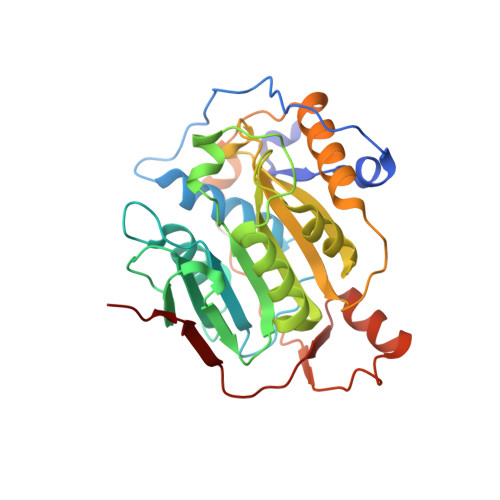
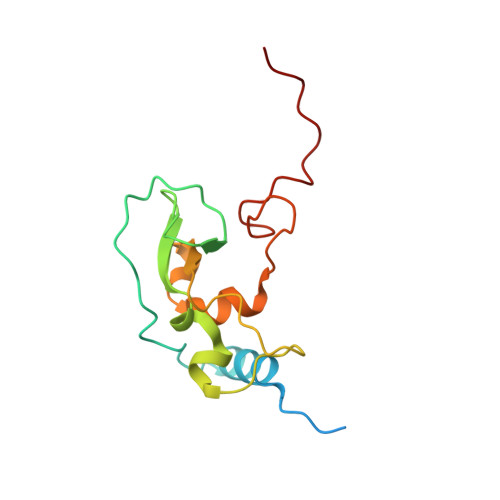
High-resolution structures of the SARS-CoV-2 2'- O -methyltransferase reveal strategies for structure-based inhibitor design.
Rosas-Lemus, M., Minasov, G., Shuvalova, L., Inniss, N.L., Kiryukhina, O., Brunzelle, J., Satchell, K.J.F.(2020) Sci Signal 13
- PubMed: 32994211 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/scisignal.abe1202
- Primary Citation Related Structures:
6W4H, 6W75, 6WJT, 6WKQ, 6WQ3, 6WRZ, 6WVN - PubMed Abstract:
There are currently no antiviral therapies specific for SARS-CoV-2, the virus responsible for the global pandemic disease COVID-19. To facilitate structure-based drug design, we conducted an x-ray crystallographic study of the SARS-CoV-2 nsp16-nsp10 2'- O -methyltransferase complex, which methylates Cap-0 viral mRNAs to improve viral protein translation and to avoid host immune detection. We determined the structures for nsp16-nsp10 heterodimers bound to the methyl donor S -adenosylmethionine (SAM), the reaction product S -adenosylhomocysteine (SAH), or the SAH analog sinefungin (SFG). We also solved structures for nsp16-nsp10 in complex with the methylated Cap-0 analog m 7 GpppA and either SAM or SAH. Comparative analyses between these structures and published structures for nsp16 from other betacoronaviruses revealed flexible loops in open and closed conformations at the m 7 GpppA-binding pocket. Bound sulfates in several of the structures suggested the location of the ribonucleic acid backbone phosphates in the ribonucleotide-binding groove. Additional nucleotide-binding sites were found on the face of the protein opposite the active site. These various sites and the conserved dimer interface could be exploited for the development of antiviral inhibitors.
- Department of Microbiology-Immunology, Northwestern University, Feinberg School of Medicine, Chicago, IL 60611, USA.
Organizational Affiliation: