Automated Design by Structure-Based Stabilization and Consensus Repair to Achieve Prefusion-Closed Envelope Trimers in a Wide Variety of HIV Strains.
Rawi, R., Rutten, L., Lai, Y.T., Olia, A.S., Blokland, S., Juraszek, J., Shen, C.H., Tsybovsky, Y., Verardi, R., Yang, Y., Zhang, B., Zhou, T., Chuang, G.Y., Kwong, P.D., Langedijk, J.P.M.(2020) Cell Rep 33: 108432-108432
- PubMed: 33238130 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.celrep.2020.108432
- Primary Citation Related Structures:
6WIX - PubMed Abstract:
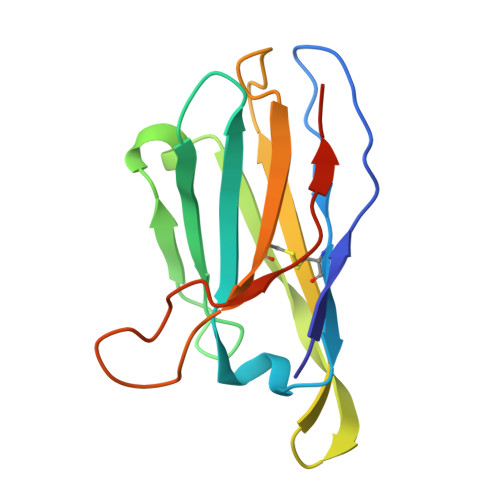
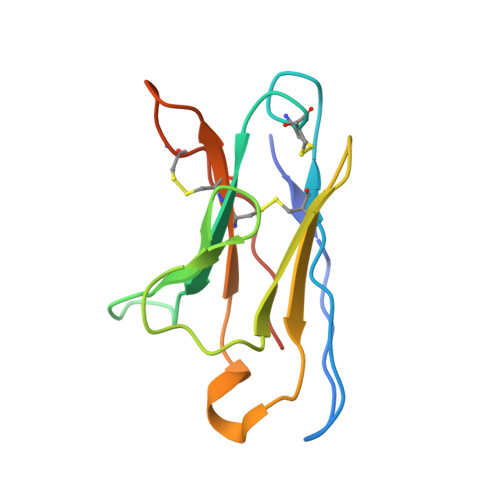
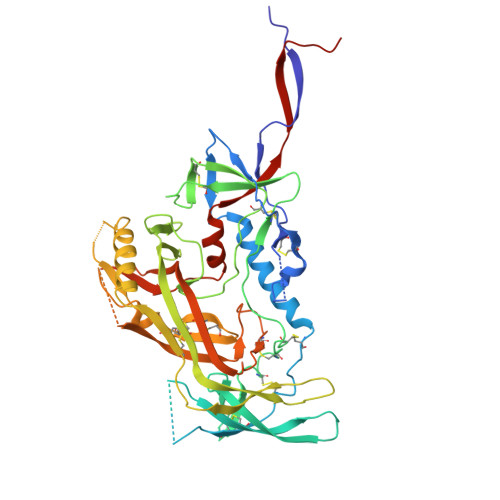
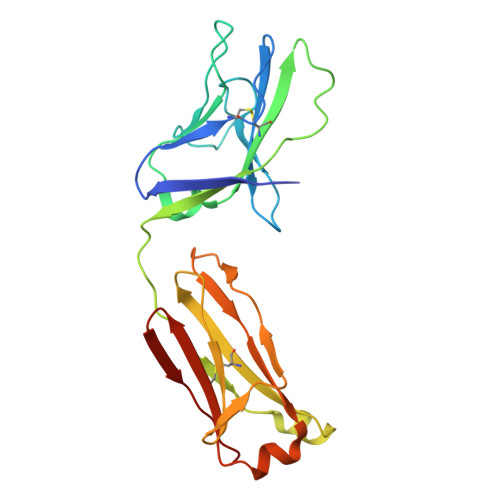
Soluble envelope (Env) trimers, stabilized in a prefusion-closed conformation, can elicit neutralizing responses against HIV-1 strains closely related to the immunizing trimer. However, to date such stabilization has succeeded with only a limited number of HIV-1 strains. To address this issue, here we develop ADROITrimer, an automated procedure involving structure-based stabilization and consensus repair, and generate "RnS-DS-SOSIP"-stabilized Envs from 180 diverse Env sequences. The vast majority of these RnS-DS-SOSIP Envs fold into prefusion-closed conformations as judged by antigenic analysis and size exclusion chromatography. Additionally, representative strains from clades AE, B, and C are stabilized in prefusion-closed conformations as shown by negative-stain electron microscopy, and the crystal structure of a clade A strain MI369.A5 Env trimer provides 3.5 Å resolution detail into stabilization and repair mutations. The automated procedure reported herein that yields well-behaved, soluble, prefusion-closed Env trimers from a majority of HIV-1 strains could have substantial impact on the development of an HIV-1 vaccine.
- Vaccine Research Center, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Bethesda, MD 20892, USA.
Organizational Affiliation: