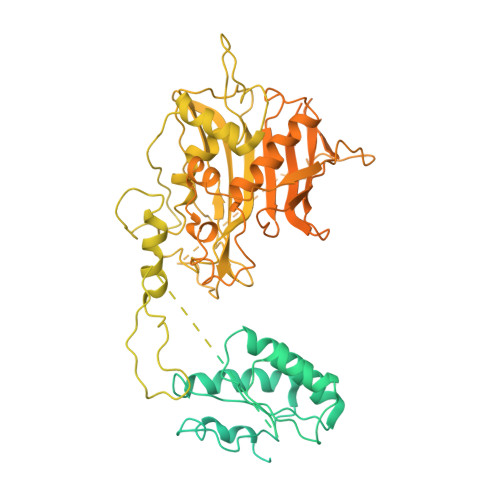
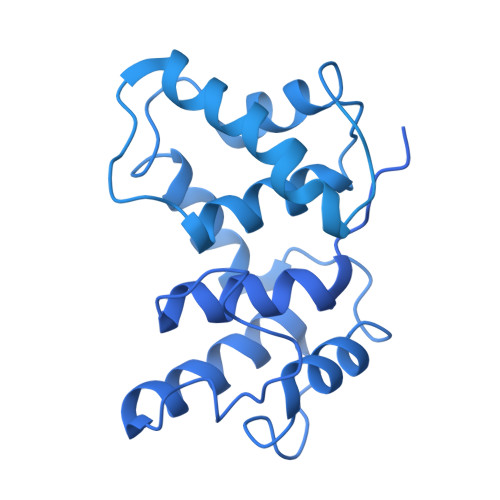
Steric occlusion regulates proximal interactions of acyl carrier protein domain in fungal fatty acid synthase.
Lou, J.W., Mazhab-Jafari, M.T.(2020) Commun Biol 3: 274-274
- PubMed: 32471977 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s42003-020-0997-y
- Primary Citation Related Structures:
6WC7 - PubMed Abstract:
The acyl carrier protein (ACP) domain shuttles substrates and reaction intermediates in type I fungal fatty acid synthases via transient protein-protein interactions. Here, using electron cryo-microscopy (cryoEM), we report the structure of a fungal FAS stalled at the dehydration reaction, which precedes the final enoyl reduction in the fatty acid biosynthesis cycle. This conformation revealed multiple contact sites between ACP and the dehydratase (DH) and enoyl reductase (ER) domains that occluded the ACP binding to the adjacent ER domain. Our data suggests a minimal path from the DH to the ER reaction site that requires minute changes in the coordinates of the structured N- and C- termini of the ACP domain.
- Department of Medical Biophysics, University of Toronto, Toronto, Canada.
Organizational Affiliation: