Ion binding properties of a naturally occurring metalloantibody
Farokhi, E., Fleming, J.K., Erasmus, M.F., Ward, A.D., Wu, Y., Gutierrez, M.G., Wojciak, J.M., Huxford, T.(2020) Antibodies (Basel) 9: 10
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
(2020) Antibodies (Basel) 9: 10
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Sonepcizumab antibody Fab fragment, heavy chain | A [auth H] | 225 | Mus musculus | Mutation(s): 0 | 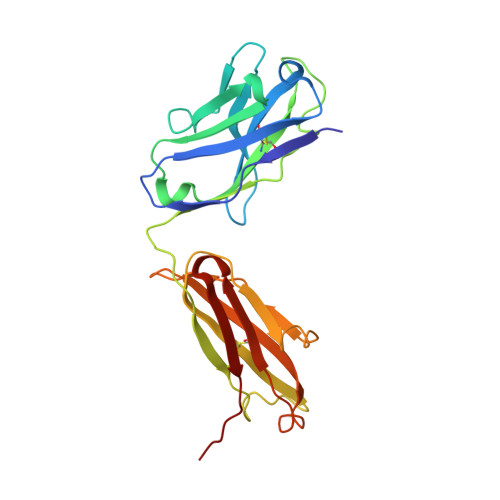 |
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Sonepcizumab antibody Fab fragment, light chain | B [auth L] | 214 | Mus musculus | Mutation(s): 0 | 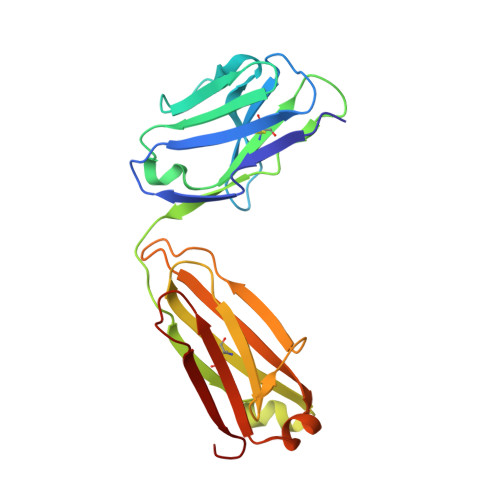 |
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| SO4 Download:Ideal Coordinates CCD File | C [auth H] | SULFATE ION O4 S QAOWNCQODCNURD-UHFFFAOYSA-L |  | ||
| CA Download:Ideal Coordinates CCD File | D [auth L], E [auth L] | CALCIUM ION Ca BHPQYMZQTOCNFJ-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 87.802 | α = 90 |
| b = 114.408 | β = 90 |
| c = 133.702 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| HKL-2000 | data reduction |
| HKL-2000 | data scaling |
| PHENIX | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Science Foundation (United States) | United States | CHE-1112547 |