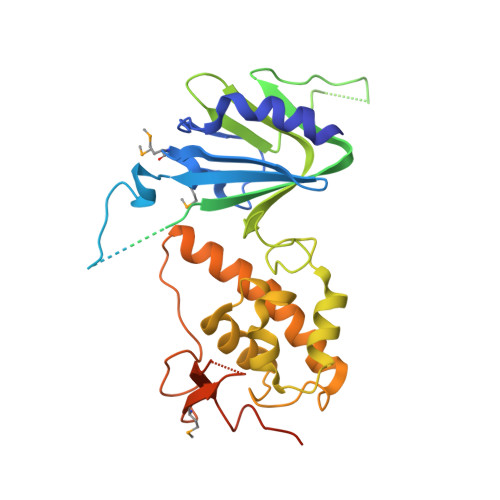
Unique Structural Features of Mammalian NEIL2 DNA Glycosylase Prime Its Activity for Diverse DNA Substrates and Environments.
Eckenroth, B.E., Cao, V.B., Averill, A.M., Dragon, J.A., Doublie, S.(2021) Structure 29: 29-42.e4
- PubMed: 32846144 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2020.08.001
- Primary Citation Related Structures:
6VJI - PubMed Abstract:
Oxidative damage on DNA arising from both endogenous and exogenous sources can result in base modifications that promote errors in replication as well as generating sites of base loss (abasic sites) that present unique challenges to maintaining genomic integrity. These lesions are excised by DNA glycosylases in the first step of the base excision repair pathway. Here we present the first crystal structure of a NEIL2 glycosylase, an enzyme active on cytosine oxidation products and abasic sites. The structure reveals an unusual "open" conformation not seen in NEIL1 or NEIL3 orthologs. NEIL2 is predicted to adopt a "closed" conformation when bound to its substrate. Combined crystallographic and solution-scattering studies show the enzyme to be conformationally dynamic in a manner distinct among the NEIL glycosylases and provide insight into the unique substrate preference of this enzyme. In addition, we characterized three cancer variants of human NEIL2, namely S140N, G230W, and G303R.
- Department of Microbiology and Molecular Genetics, University of Vermont, Stafford Hall, 95 Carrigan Drive, Burlington, VT 05405, USA. Electronic address: beckenro@uvm.edu.
Organizational Affiliation: