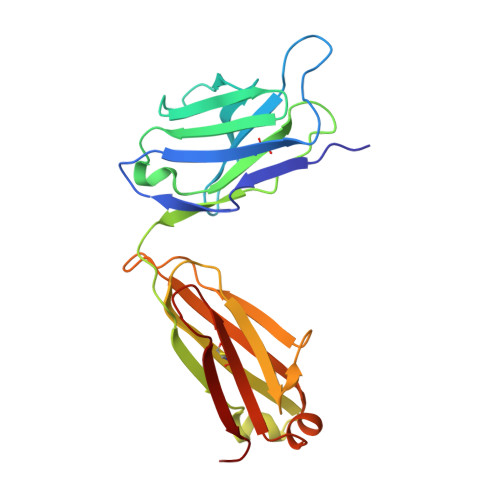
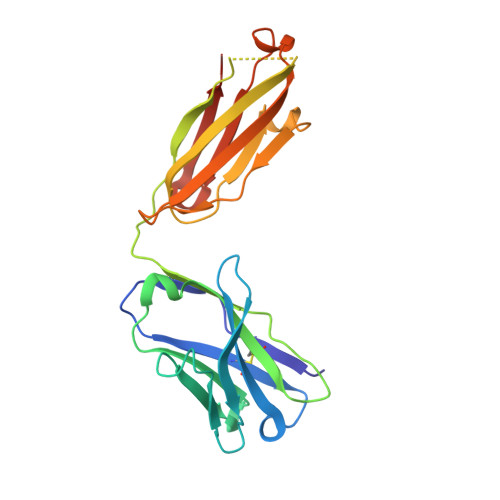
HIV vaccine delayed boosting increases Env variable region 2-specific antibody effector functions.
Easterhoff, D., Pollara, J., Luo, K., Janus, B., Gohain, N., Williams, L.D., Tay, M.Z., Monroe, A., Peachman, K., Choe, M., Min, S., Lusso, P., Zhang, P., Go, E.P., Desaire, H., Bonsignori, M., Hwang, K.K., Beck, C., Kakalis, M., O'Connell, R.J., Vasan, S., Kim, J.H., Michael, N.L., Excler, J.L., Robb, M.L., Rerks-Ngarm, S., Kaewkungwal, J., Pitisuttithum, P., Nitayaphan, S., Sinangil, F., Tartaglia, J., Phogat, S., Wiehe, K., Saunders, K.O., Montefiori, D.C., Tomaras, G.D., Moody, M.A., Arthos, J., Rao, M., Joyce, M.G., Ofek, G.A., Ferrari, G., Haynes, B.F.(2020) JCI Insight 5
- PubMed: 31996483 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1172/jci.insight.131437
- Primary Citation Related Structures:
6VBO, 6VBP, 6VBQ - PubMed Abstract:
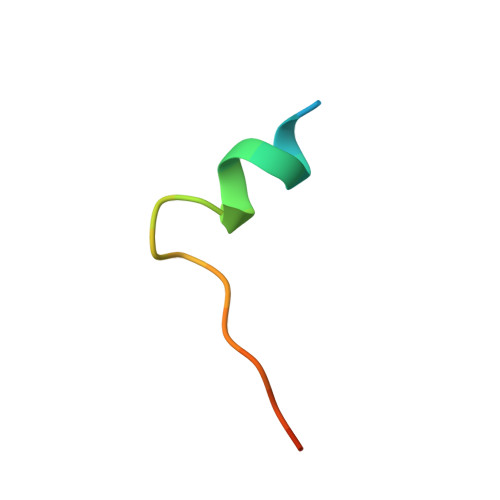
In the RV144 HIV-1 phase III trial, vaccine efficacy directly correlated with the magnitude of the variable region 2-specific (V2-specific) IgG antibody response, and in the presence of low plasma IgA levels, with the magnitude of plasma antibody-dependent cellular cytotoxicity. Reenrollment of RV144 vaccinees in the RV305 trial offered the opportunity to define the function, maturation, and persistence of vaccine-induced V2-specific and other mAb responses after boosting. We show that the RV144 vaccine regimen induced persistent V2 and other HIV-1 envelope-specific memory B cell clonal lineages that could be identified throughout the approximately 11-year vaccination period. Subsequent boosts increased somatic hypermutation, a critical requirement for antibody affinity maturation. Characterization of 22 vaccine-induced V2-specific mAbs with epitope specificities distinct from previously characterized RV144 V2-specific mAbs CH58 and CH59 found increased in vitro antibody-mediated effector functions. Thus, when inducing non-neutralizing antibodies, one method by which to improve HIV-1 vaccine efficacy may be through late boosting to diversify the V2-specific response to increase the breadth of antibody-mediated anti-HIV-1 effector functions.
- Duke Human Vaccine Institute, Duke University, Durham, North Carolina, USA.
Organizational Affiliation: