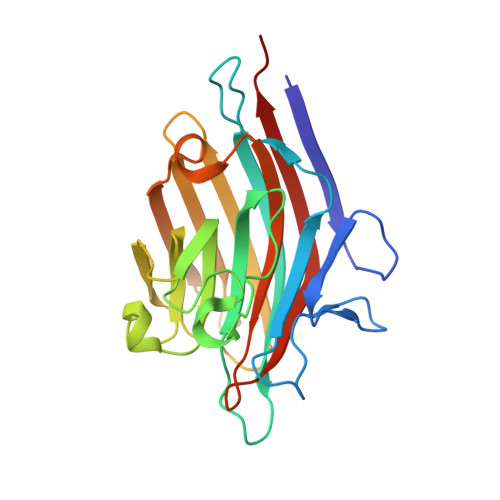
Crystal structures of peanut lectin in the presence of synthetic beta-N- and beta-S-galactosides disclose evidence for the recognition of different glycomimetic ligands.
Cagnoni, A.J., Primo, E.D., Klinke, S., Cano, M.E., Giordano, W., Marino, K.V., Kovensky, J., Goldbaum, F.A., Uhrig, M.L., Otero, L.H.(2020) Acta Crystallogr D Struct Biol 76: 1080-1091
- PubMed: 33135679 Search on PubMed
- DOI: https://doi.org/10.1107/S2059798320012371
- Primary Citation Related Structures:
6V95, 6VAV, 6VAW, 6VC3, 6VC4, 6VGF - PubMed Abstract:
Carbohydrate-lectin interactions are involved in important cellular recognition processes, including viral and bacterial infections, inflammation and tumor metastasis. Hence, structural studies of lectin-synthetic glycan complexes are essential for understanding lectin-recognition processes and for the further design of promising chemotherapeutics that interfere with sugar-lectin interactions. Plant lectins are excellent models for the study of the molecular-recognition process. Among them, peanut lectin (PNA) is highly relevant in the field of glycobiology because of its specificity for β-galactosides, showing high affinity towards the Thomsen-Friedenreich antigen, a well known tumor-associated carbohydrate antigen. Given this specificity, PNA is one of the most frequently used molecular probes for the recognition of tumor cell-surface O-glycans. Thus, it has been extensively used in glycobiology for inhibition studies with a variety of β-galactoside and β-lactoside ligands. Here, crystal structures of PNA are reported in complex with six novel synthetic hydrolytically stable β-N- and β-S-galactosides. These complexes disclosed key molecular-binding interactions of the different sugars with PNA at the atomic level, revealing the roles of specific water molecules in protein-ligand recognition. Furthermore, binding-affinity studies by isothermal titration calorimetry showed dissociation-constant values in the micromolar range, as well as a positive multivalency effect in terms of affinity in the case of the divalent compounds. Taken together, this work provides a qualitative structural rationale for the upcoming synthesis of optimized glycoclusters designed for the study of lectin-mediated biological processes. The understanding of the recognition of β-N- and β-S-galactosides by PNA represents a benchmark in protein-carbohydrate interactions since they are novel synthetic ligands that do not belong to the family of O-linked glycosides.
- Laboratorio de Glicómica Funcional y Molecular, Instituto de Biología y Medicina Experimental, IBYME-CONICET, Vuelta de Obligado 2490, C1428ADN Buenos Aires, Argentina.
Organizational Affiliation: