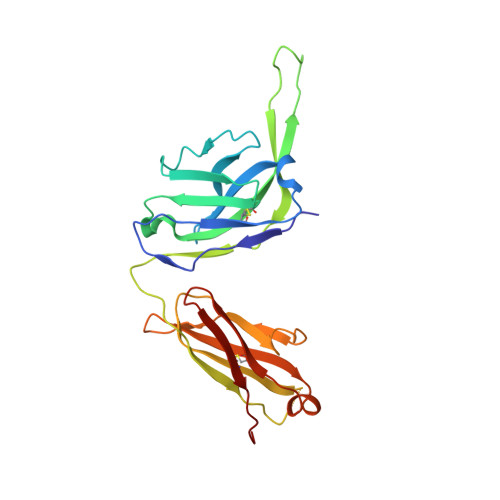
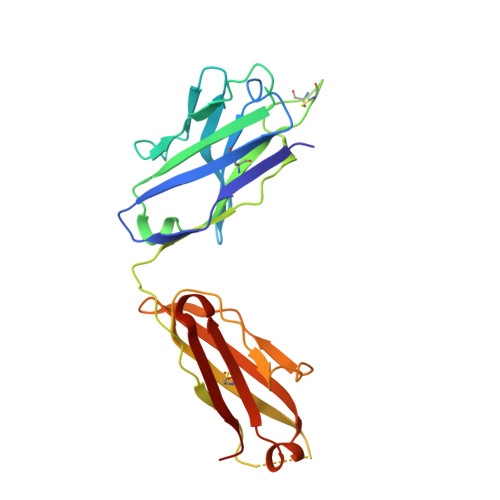
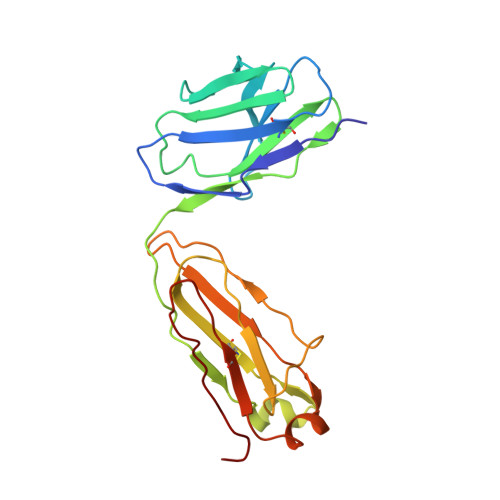
Disruption of the HIV-1 Envelope allosteric network blocks CD4-induced rearrangements.
Henderson, R., Lu, M., Zhou, Y., Mu, Z., Parks, R., Han, Q., Hsu, A.L., Carter, E., Blanchard, S.C., Edwards, R.J., Wiehe, K., Saunders, K.O., Borgnia, M.J., Bartesaghi, A., Mothes, W., Haynes, B.F., Acharya, P., Munir Alam, S.(2020) Nat Commun 11: 520-520
- PubMed: 31980614 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-019-14196-w
- Primary Citation Related Structures:
6V8X, 6V8Z - PubMed Abstract:
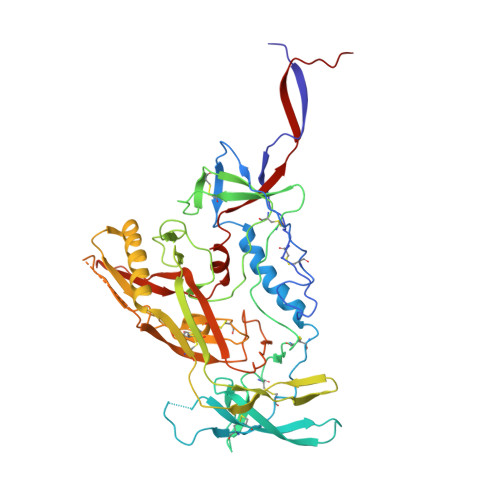
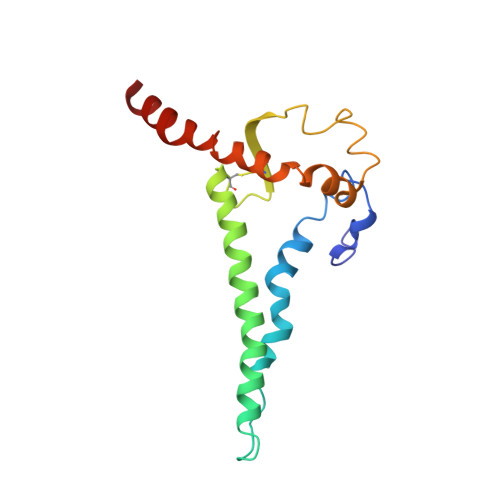
The trimeric HIV-1 Envelope protein (Env) mediates viral-host cell fusion via a network of conformational transitions, with allosteric elements in each protomer orchestrating host receptor-induced exposure of the co-receptor binding site and fusion elements. To understand the molecular details of this allostery, here, we introduce Env mutations aimed to prevent CD4-induced rearrangements in the HIV-1 BG505 Env trimer. Binding analysis and single-molecule Förster Resonance Energy Transfer confirm that these mutations prevent CD4-induced transitions of the HIV-1 Env. Structural analysis by single-particle cryo-electron microscopy performed on the BG505 SOSIP mutant Env proteins shows rearrangements in the gp120 topological layer contacts with gp41. Displacement of a conserved tryptophan (W571) from its typical pocket in these Env mutants renders the Env insensitive to CD4 binding. These results reveal the critical function of W571 as a conformational switch in Env allostery and receptor-mediated viral entry and provide insights on Env conformation that are relevant for vaccine design.
- Department of Medicine, Duke University School of Medicine, Durham, NC, 27710, USA. rory.henderson@duke.edu.
Organizational Affiliation: