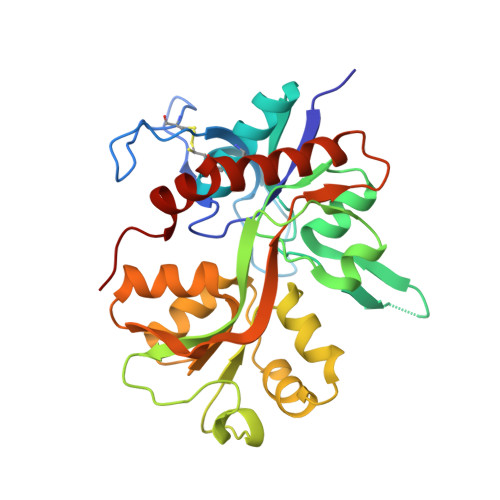
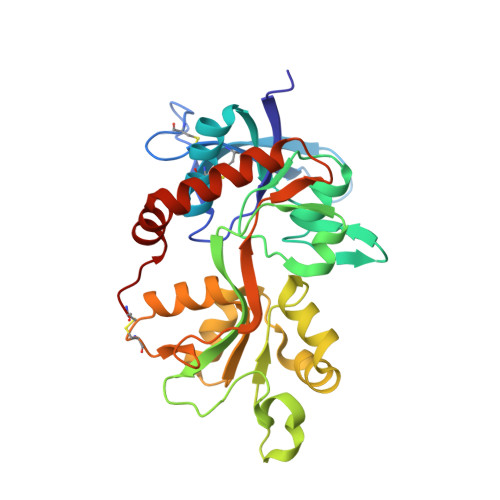
Structural basis of subtype-selective competitive antagonism for GluN2C/2D-containing NMDA receptors.
Wang, J.X., Irvine, M.W., Burnell, E.S., Sapkota, K., Thatcher, R.J., Li, M., Simorowski, N., Volianskis, A., Collingridge, G.L., Monaghan, D.T., Jane, D.E., Furukawa, H.(2020) Nat Commun 11: 423-423
- PubMed: 31969570 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-020-14321-0
- Primary Citation Related Structures:
6UZ6, 6UZG, 6UZR, 6UZW, 6UZX - PubMed Abstract:
N-Methyl-D-aspartate receptors (NMDARs) play critical roles in the central nervous system. Their heterotetrameric composition generates subtypes with distinct functional properties and spatio-temporal distribution in the brain, raising the possibility for subtype-specific targeting by pharmacological means for treatment of neurological diseases. While specific compounds for GluN2A and GluN2B-containing NMDARs are well established, those that target GluN2C and GluN2D are currently underdeveloped with low potency and uncharacterized binding modes. Here, using electrophysiology and X-ray crystallography, we show that UBP791 ((2S*,3R*)-1-(7-(2-carboxyethyl)phenanthrene-2-carbonyl)piperazine-2,3-dicarboxylic acid) inhibits GluN2C/2D with 40-fold selectivity over GluN2A-containing receptors, and that a methionine and a lysine residue in the ligand binding pocket (GluN2D-Met763/Lys766, GluN2C-Met736/Lys739) are the critical molecular elements for the subtype-specific binding. These findings led to development of UBP1700 ((2S*,3R*)-1-(7-(2-carboxyvinyl)phenanthrene-2-carbonyl)piperazine-2,3-dicarboxylic acid) which shows over 50-fold GluN2C/2D-selectivity over GluN2A with potencies in the low nanomolar range. Our study shows that the L-glutamate binding site can be targeted for GluN2C/2D-specific inhibition.
- WM Keck Structural Biology Laboratory, Cold Spring Harbor Laboratory, Cold Spring Harbor, NY, 11724, USA.
Organizational Affiliation: