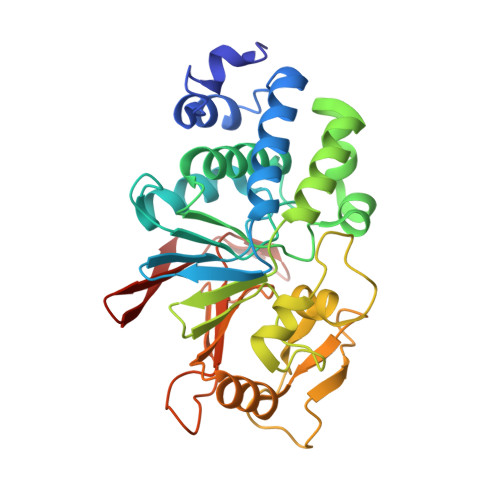
The structure of the RCAN1:CN complex explains the inhibition of and substrate recruitment by calcineurin.
Li, Y., Sheftic, S.R., Grigoriu, S., Schwieters, C.D., Page, R., Peti, W.(2020) Sci Adv 6
- PubMed: 32936779 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.aba3681
- Primary Citation Related Structures:
6UUQ - PubMed Abstract:
Regulator of calcineurin 1 (RCAN1) is an endogenous inhibitor of the Ser/Thr phosphatase calcineurin (CN). It has been shown that excessive inhibition of CN is a critical factor for Down syndrome and Alzheimer's disease. Here, we determined RCAN1's mode of action. Using a combination of structural, biophysical, and biochemical studies, we show that RCAN1 inhibits CN via multiple routes: first, by blocking essential substrate recruitment sites and, second, by blocking the CN active site using two distinct mechanisms. We also show that phosphorylation either inhibits RCAN1-CN assembly or converts RCAN1 into a weak inhibitor, which can be reversed by CN via dephosphorylation. This highlights the interplay between posttranslational modifications in regulating CN activity. Last, this work advances our understanding of how active site inhibition of CN can be achieved in a highly specific manner. Together, these data provide the necessary road map for targeting multiple neurological disorders.
- Department of Chemistry and Biochemistry, University of Arizona, 1041 E. Lowell St., Tucson, AZ 85721, USA.
Organizational Affiliation: