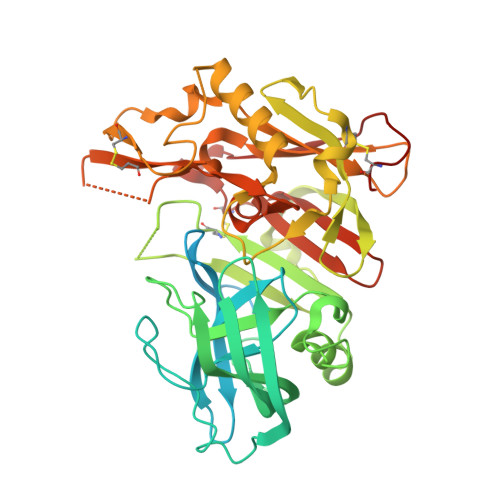
A Structure-Based Discovery Platform for BACE2 and the Development of Selective BACE Inhibitors.
Yen, Y.C., Kammeyer, A.M., Tirlangi, J., Ghosh, A.K., Mesecar, A.D.(2021) ACS Chem Neurosci 12: 581-588
- PubMed: 33544569 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acschemneuro.0c00629
- Primary Citation Related Structures:
6UJ0, 6UJ1 - PubMed Abstract:
The ability to perform routine structure-guided drug design for selective BACE inhibitors has been limited because of the lack of robust platform for BACE2 expression, purification, and crystallization. To overcome this limitation, we developed a platform that produces 2-3 mg of pure BACE2 protein per liter of E. coli culture, and we used this protein to design macrocyclic compounds that potently and selectively inhibit BACE1 over BACE2. Compound 2 was found to potently inhibit BACE 1 ( K i = 5 nM) with a selectivity of 214-fold over BACE2. The X-ray crystal structures of unbound BACE2 (2.2 Å) and BACE2 bound to compound 3 (3.0 Å and K i = 7 nM) were determined and compared to the X-ray structures of BACE1 revealing the S1-S3 subsite as a selectivity determinant. This platform should enable a more rapid development of new and selective BACE inhibitors for the treatment of Alzheimer's disease or type II diabetes.