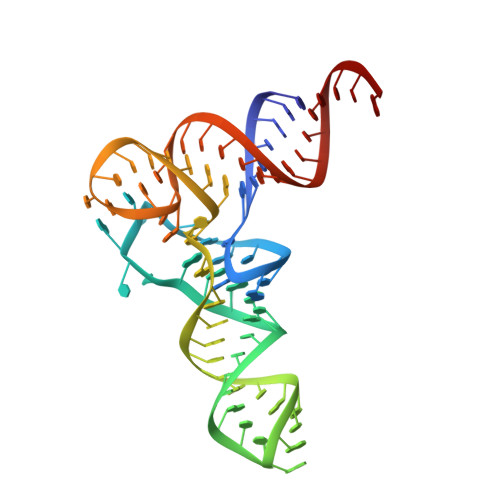
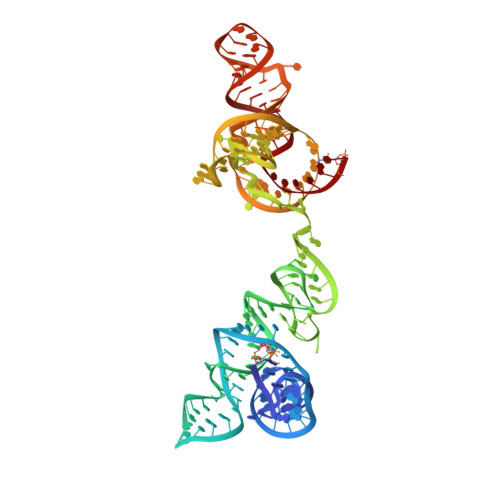Structural basis for tRNA decoding and aminoacylation sensing by T-box riboregulators.
Battaglia, R.A., Grigg, J.C., Ke, A.(2019) Nat Struct Mol Biol 26: 1106-1113
- PubMed: 31740853 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-019-0327-6
- Primary Citation Related Structures:
6UFG, 6UFH - PubMed Abstract:
T-box riboregulators are a class of cis-regulatory RNAs that govern the bacterial response to amino acid starvation by binding, decoding and reading the aminoacylation status of specific transfer RNAs. Here we provide a high-resolution crystal structure of a full-length T-box from Mycobacterium tuberculosis that explains tRNA decoding and aminoacylation sensing by this riboregulator. Overall, the T-box consists of decoding and aminoacylation sensing modules bridged by a rigid pseudoknot structure formed by the mid-region domains. Stem-I and the Stem-II S-turn assemble a claw-like decoding module, while the antiterminator, Stem-III, and the adjacent linker form a tightly interwoven aminoacylation sensing module. The uncharged tRNA is selectively recognized by an unexpected set of favorable contacts from the linker region in the aminoacylation sensing module. A complex structure with a charged tRNA mimic shows that the extra moiety dislodges the linker, which is indicative of the possible chain of events that lead to alternative base-pairing and altered expression output.
- Department of Molecular Biology and Genetics, Ithaca, NY, USA.
Organizational Affiliation: