Structural and genetic convergence of HIV-1 neutralizing antibodies in vaccinated non-human primates.
Cai, F., Chen, W.H., Wu, W., Jones, J.A., Choe, M., Gohain, N., Shen, X., LaBranche, C., Eaton, A., Sutherland, L., Lee, E.M., Hernandez, G.E., Wu, N.R., Scearce, R., Seaman, M.S., Moody, M.A., Santra, S., Wiehe, K., Tomaras, G.D., Wagh, K., Korber, B., Bonsignori, M., Montefiori, D.C., Haynes, B.F., de Val, N., Joyce, M.G., Saunders, K.O.(2021) PLoS Pathog 17: e1009624-e1009624
- PubMed: 34086838 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1009624
- Primary Citation Related Structures:
6U6M, 6U6O - PubMed Abstract:
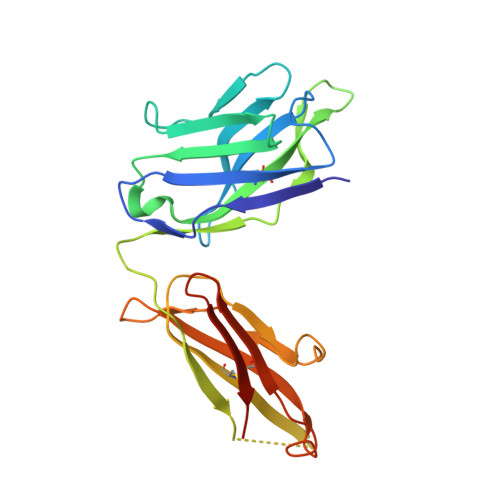
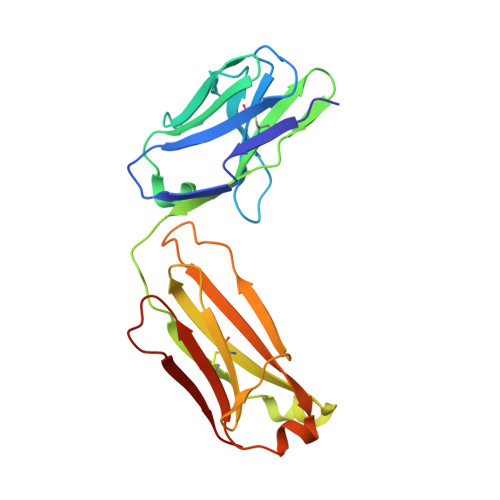
A primary goal of HIV-1 vaccine development is the consistent elicitation of protective, neutralizing antibodies. While highly similar neutralizing antibodies (nAbs) have been isolated from multiple HIV-infected individuals, it is unclear whether vaccination can consistently elicit highly similar nAbs in genetically diverse primates. Here, we show in three outbred rhesus macaques that immunization with Env elicits a genotypically and phenotypically conserved nAb response. From these vaccinated macaques, we isolated four antibody lineages that had commonalities in immunoglobulin variable, diversity, and joining gene segment usage. Atomic-level structures of the antigen binding fragments of the two most similar antibodies showed nearly identical paratopes. The Env binding modes of each of the four vaccine-induced nAbs were distinct from previously known monoclonal HIV-1 neutralizing antibodies, but were nearly identical to each other. The similarities of these antibodies show that the immune system in outbred primates can respond to HIV-1 Env vaccination with a similar structural and genotypic solution for recognizing a particular neutralizing epitope. These results support rational vaccine design for HIV-1 that aims to reproducibly elicit, in genetically diverse primates, nAbs with specific paratope structures capable of binding conserved epitopes.
- Duke Human Vaccine Institute, Duke University Medical Center, Durham, North Carolina, United States of America.
Organizational Affiliation: