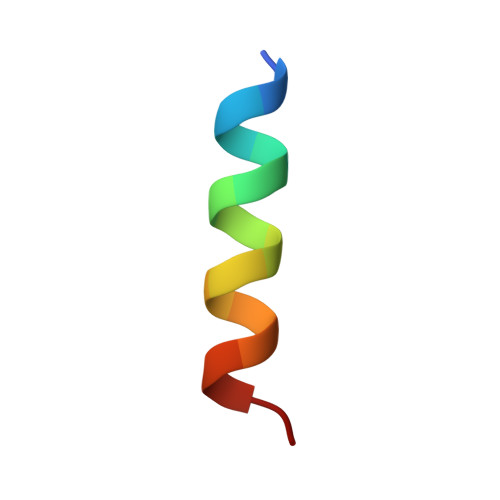
Talin mechanosensitivity is modulated by a direct interaction with cyclin-dependent kinase-1.
Gough, R.E., Jones, M.C., Zacharchenko, T., Le, S., Yu, M., Jacquemet, G., Muench, S.P., Yan, J., Humphries, J.D., Jorgensen, C., Humphries, M.J., Goult, B.T.(2021) J Biological Chem 297: 100837-100837
- PubMed: 34118235 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbc.2021.100837
- Primary Citation Related Structures:
6TWN - PubMed Abstract:
Talin (TLN1) is a mechanosensitive component of adhesion complexes that directly couples integrins to the actin cytoskeleton. In response to force, talin undergoes switch-like behavior of its multiple rod domains that modulate interactions with its binding partners. Cyclin-dependent kinase-1 (CDK1) is a key regulator of the cell cycle, exerting its effects through synchronized phosphorylation of a large number of protein targets. CDK1 activity maintains adhesion during interphase, and its inhibition is a prerequisite for the tightly choreographed changes in cell shape and adhesion that are required for successful mitosis. Using a combination of biochemical, structural, and cell biological approaches, we demonstrate a direct interaction between talin and CDK1 that occurs at sites of integrin-mediated adhesion. Mutagenesis demonstrated that CDK1 contains a functional talin-binding LD motif, and the binding site within talin was pinpointed to helical bundle R8. Talin also contains a consensus CDK1 phosphorylation motif centered on S1589, a site shown to be phosphorylated by CDK1 in vitro. A phosphomimetic mutant of this site within talin lowered the binding affinity of the cytoskeletal adaptor KANK and weakened the response of this region to force as measured by single molecule stretching, potentially altering downstream mechanotransduction pathways. The direct binding of the master cell cycle regulator CDK1 to the primary integrin effector talin represents a coupling of cell proliferation and cell adhesion machineries and thereby indicates a mechanism by which the microenvironment can control cell division in multicellular organisms.
- School of Biosciences, University of Kent, Canterbury, Kent, UK.
Organizational Affiliation: