Plasmodium vivax N-myristoyltransferase with bound indazole inhibitor IMP-917
Brannigan, J.A.To be published.
Experimental Data Snapshot
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Glycylpeptide N-tetradecanoyltransferase | A [auth AAA], B [auth BBB], C [auth CCC] | 388 | Plasmodium vivax | Mutation(s): 0 Gene Names: PVX_085815 EC: 2.3.1.97 | 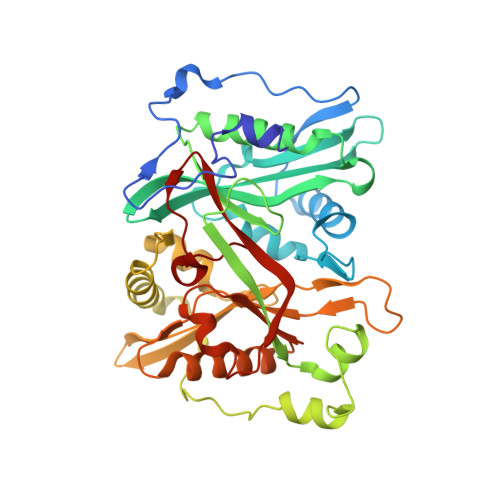 |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | A5K1A2 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 6 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| MYA Download:Ideal Coordinates CCD File | E [auth AAA], L [auth BBB], Q [auth CCC] | TETRADECANOYL-COA C35 H62 N7 O17 P3 S DUAFKXOFBZQTQE-QSGBVPJFSA-N |  | ||
| 9M2 (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | D [auth AAA], K [auth BBB], P [auth CCC] | 1-[5-[4-fluoranyl-2-[2-(1,3,5-trimethylpyrazol-4-yl)ethoxy]phenyl]-2~{H}-indazol-3-yl]-~{N},~{N}-dimethyl-methanamine C24 H28 F N5 O CGEINWMYWNHNPM-UHFFFAOYSA-N |  | ||
| SO4 Download:Ideal Coordinates CCD File | I [auth AAA], J [auth AAA], U [auth CCC] | SULFATE ION O4 S QAOWNCQODCNURD-UHFFFAOYSA-L |  | ||
| DMS Download:Ideal Coordinates CCD File | G [auth AAA], N [auth BBB], S [auth CCC] | DIMETHYL SULFOXIDE C2 H6 O S IAZDPXIOMUYVGZ-UHFFFAOYSA-N |  | ||
| CL Download:Ideal Coordinates CCD File | H [auth AAA], O [auth BBB], T [auth CCC] | CHLORIDE ION Cl VEXZGXHMUGYJMC-UHFFFAOYSA-M |  | ||
| MG Download:Ideal Coordinates CCD File | F [auth AAA], M [auth BBB], R [auth CCC] | MAGNESIUM ION Mg JLVVSXFLKOJNIY-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 57.472 | α = 90 |
| b = 121.655 | β = 90 |
| c = 178.371 | γ = 90 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| XDS | data reduction |
| SCALA | data scaling |
| REFMAC | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Wellcome Trust | United Kingdom | -- |