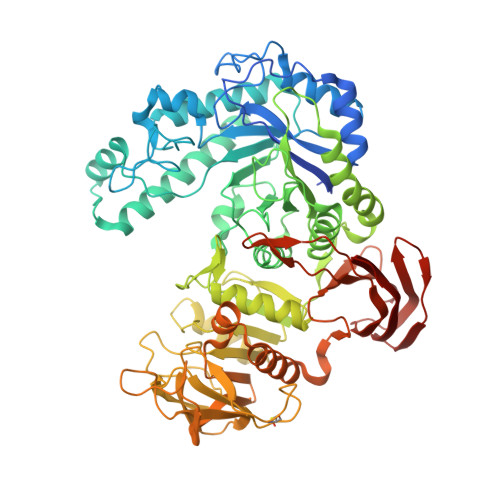
The first structure-function study of GH151 alpha-l-fucosidase uncovers new oligomerization pattern, active site complementation, and selective substrate specificity
Koval'ova, T., Koval, T., Stransky, J., Kolenko, P., Duskova, J., Svecova, L., Vodickova, P., Spiwok, V., Benesova, E., Lipovova, P., Dohnalek, J.(2022) FEBS J