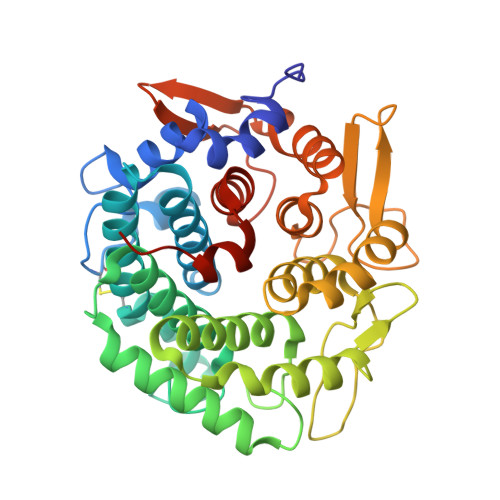
Mutational Analysis of Linalool Dehydratase Isomerase Suggests That Alcohol and Alkene Transformations Are Catalyzed Using Noncovalent Mechanisms
Cuetos, A., Iglesias-Fernandez, J., Danesh-Azari, H.R., Zukic, E., Dowle, A., Osuna, S., Grogan, G.(2020) ACS Catal