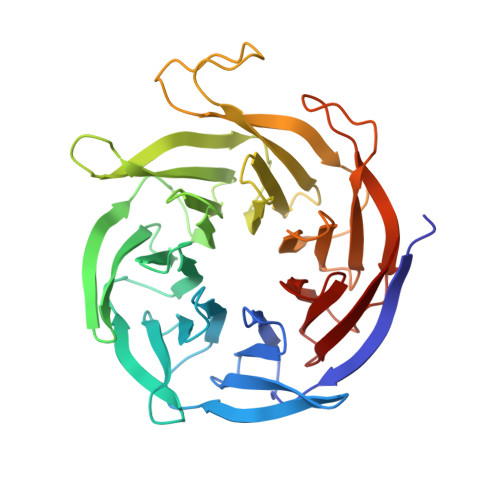
Structural insights into Fe-S protein biogenesis by the CIA targeting complex.
Kassube, S.A., Thoma, N.H.(2020) Nat Struct Mol Biol 27: 735-742
- PubMed: 32632277 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-020-0454-0
- Primary Citation Related Structures:
6TBL, 6TBN, 6TC0 - PubMed Abstract:
The cytosolic iron-sulfur (Fe-S) assembly (CIA) pathway is required for the insertion of Fe-S clusters into cytosolic and nuclear client proteins, including many DNA replication and repair factors. The molecular mechanisms of client protein recognition and Fe-S cluster transfer remain unknown. Here, we report crystal structures of the CIA targeting complex (CTC), revealing that its CIAO2B subunit is centrally located and bridges CIAO1 and the client adaptor protein MMS19. Cryo-EM reconstructions of human CTC bound either to the DNA replication factor primase or to the DNA helicase DNA2, combined with biochemical, biophysical and yeast complementation assays, reveal an evolutionarily conserved, bipartite client recognition mode facilitated by CIAO1 and the structural flexibility of the MMS19 subunit. Unexpectedly, the primase Fe-S cluster is located ~70 Å away from the CTC reactive cysteine, implicating conformational dynamics of the CTC or additional maturation factors in the mechanism of Fe-S cluster transfer.
- Friedrich Miescher Institute for Biomedical Research, Basel, Switzerland.
Organizational Affiliation: