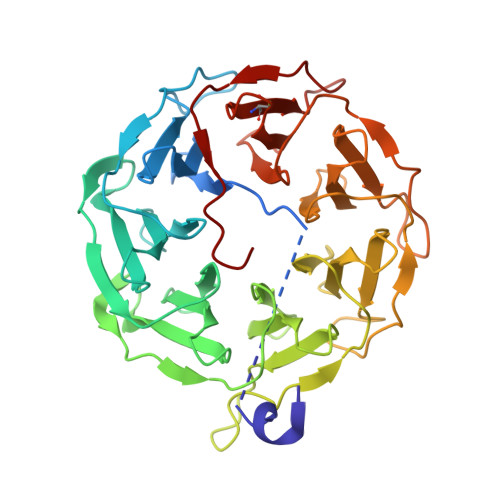
Lectin PLL3, a Novel Monomeric Member of the Seven-Bladed beta-Propeller Lectin Family.
Faltinek, L., Fujdiarova, E., Melicher, F., Houser, J., Kasakova, M., Kondakov, N., Kononov, L., Parkan, K., Vidal, S., Wimmerova, M.(2019) Molecules 24
- PubMed: 31835851 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/molecules24244540
- Primary Citation Related Structures:
6T96 - PubMed Abstract:
The Photorhabdus species is a Gram-negative bacteria of the family Morganellaceae that is known for its mutualistic relationship with Heterorhabditis nematodes and pathogenicity toward insects. This study is focused on the characterization of the recombinant lectin PLL3 with an origin in P. laumondii subsp. laumondii . PLL3 belongs to the PLL family of lectins with a seven-bladed β-propeller fold. The binding properties of PLL3 were tested by hemagglutination assay, glycan array, isothermal titration calorimetry, and surface plasmon resonance, and its structure was determined by X-ray crystallography. Obtained data revealed that PLL3 binds similar carbohydrates to those that the other PLL family members bind, with some differences in the binding properties. PLL3 exhibited the highest affinity toward l-fucose and its derivatives but was also able to interact with O -methylated glycans and other ligands. Unlike the other members of this family, PLL3 was discovered to be a monomer, which might correspond to a weaker avidity effect compared to homologous lectins. Based on the similarity to the related lectins and their proposed biological function, PLL3 might accompany them during the interaction of P. laumondii with both the nematode partner and the insect host.
- Department of Biochemistry, Faculty of Science, Masaryk University, Kotlářská 2, 611 37 Brno, Czech Republic.
Organizational Affiliation: