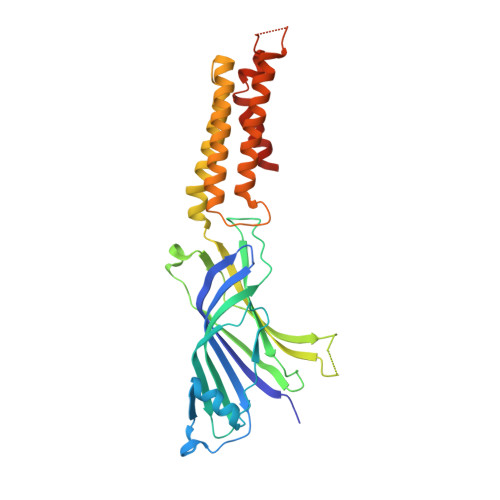
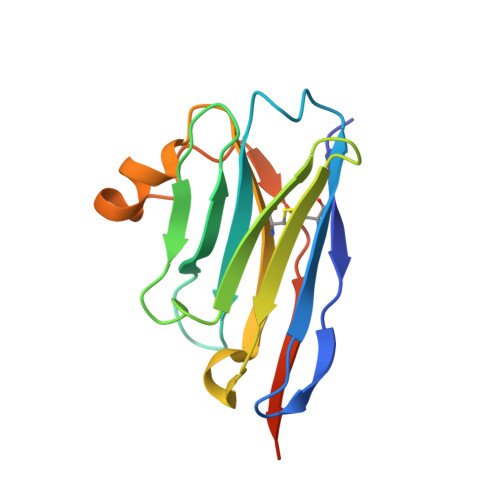
Modulation of the Erwinia ligand-gated ion channel (ELIC) and the 5-HT 3 receptor via a common vestibule site.
Brams, M., Govaerts, C., Kambara, K., Price, K.L., Spurny, R., Gharpure, A., Pardon, E., Evans, G.L., Bertrand, D., Lummis, S.C., Hibbs, R.E., Steyaert, J., Ulens, C.(2020) Elife 9
- PubMed: 31990273 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.51511
- Primary Citation Related Structures:
6SSI, 6SSP - PubMed Abstract:
Pentameric ligand-gated ion channels (pLGICs) or Cys-loop receptors are involved in fast synaptic signaling in the nervous system. Allosteric modulators bind to sites that are remote from the neurotransmitter binding site, but modify coupling of ligand binding to channel opening. In this study, we developed nanobodies (single domain antibodies), which are functionally active as allosteric modulators, and solved co-crystal structures of the prokaryote ( Erwinia ) channel ELIC bound either to a positive or a negative allosteric modulator. The allosteric nanobody binding sites partially overlap with those of small molecule modulators, including a vestibule binding site that is not accessible in some pLGICs. Using mutagenesis, we extrapolate the functional importance of the vestibule binding site to the human 5-HT 3 receptor, suggesting a common mechanism of modulation in this protein and ELIC. Thus we identify key elements of allosteric binding sites, and extend drug design possibilities in pLGICs with an accessible vestibule site.
- Laboratory of Structural Neurobiology, Department of Cellular and Molecular Medicine, Faculty of Medicine, KU Leuven, Leuven, Belgium.
Organizational Affiliation: