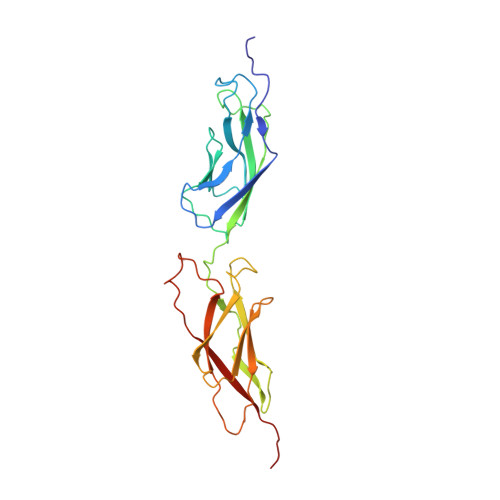
Intermediate-resolution crystal structure of the human adenovirus B serotype 3 fibre knob in complex with the EC2-EC3 fragment of desmoglein 2.
Vassal-Stermann, E., Hutin, S., Fender, P., Burmeister, W.P.(2019) Acta Crystallogr F Struct Biol Commun 75: 750-757
- PubMed: 31797817 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X19015784
- Primary Citation Related Structures:
6SIT - PubMed Abstract:
The cryo-electron microscopy (cryo-EM) structure of the complex between the trimeric human adenovirus B serotype 3 fibre knob and human desmoglein 2 fragments containing cadherin domains EC2 and EC3 has been published, showing 3:1 and 3:2 complexes. Here, the crystal structure determined at 4.5 Å resolution is presented with one EC2-EC3 desmoglein fragment bound per fibre knob monomer in the asymmetric unit, leading to an apparent 3:3 stoichiometry. However, in concentrated solution the 3:2 complex is predominant, as shown by small-angle X-ray scattering (SAXS), while cryo-EM at lower concentrations showed a majority of the 3:1 complex. Substitution of the calcium ions bound to the desmoglein domains by terbium ions allowed confirmation of the X-ray model using their anomalous scattering and shows that at least one binding site per cluster of calcium ions is intact and exchangeable and, combined with SAXS data, that the cadherin domains are folded even in the distal part that is invisible in the cryo-EM reconstruction.
- Institut de Biologie Structurale (IBS), Université Grenoble Alpes, CNRS, CEA, 71 Avenue des Martyrs, 38000 Grenoble, France.
Organizational Affiliation: