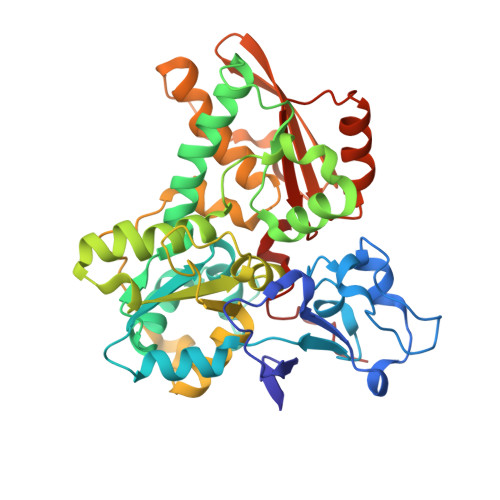
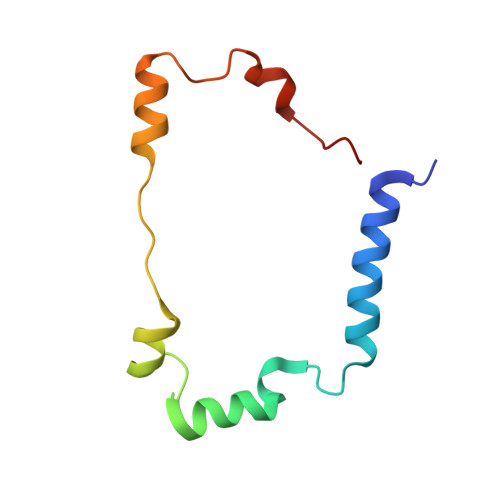
Caught in the H inact : Crystal Structure and Spectroscopy Reveal a Sulfur Bound to the Active Site of an O 2 -stable State of [FeFe] Hydrogenase.
Rodriguez-Macia, P., Galle, L.M., Bjornsson, R., Lorent, C., Zebger, I., Yoda, Y., Cramer, S.P., DeBeer, S., Span, I., Birrell, J.A.(2020) Angew Chem Int Ed Engl 59: 16786-16794
- PubMed: 32488975 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/anie.202005208
- Primary Citation Related Structures:
6SG2 - PubMed Abstract:
[FeFe] hydrogenases are the most active H 2 converting catalysts in nature, but their extreme oxygen sensitivity limits their use in technological applications. The [FeFe] hydrogenases from sulfate reducing bacteria can be purified in an O 2 -stable state called H inact . To date, the structure and mechanism of formation of H inact remain unknown. Our 1.65 Å crystal structure of this state reveals a sulfur ligand bound to the open coordination site. Furthermore, in-depth spectroscopic characterization by X-ray absorption spectroscopy (XAS), nuclear resonance vibrational spectroscopy (NRVS), resonance Raman (RR) spectroscopy and infrared (IR) spectroscopy, together with hybrid quantum mechanical and molecular mechanical (QM/MM) calculations, provide detailed chemical insight into the H inact state and its mechanism of formation. This may facilitate the design of O 2 -stable hydrogenases and molecular catalysts.
- Department of Inorganic Spectroscopy, Max Planck Institute for Chemical Energy Conversion, Stiftstraße 34-36, 45470, Mülheim an der Ruhr, Germany.
Organizational Affiliation: