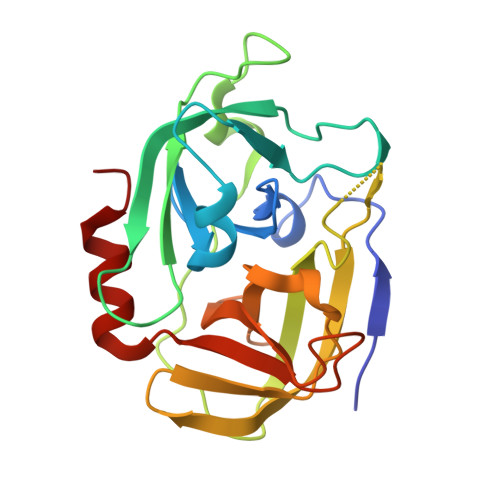
Structural Determinants of Substrate Specificity of SplF Protease from Staphylococcus aureus .
Stach, N., Karim, A., Golik, P., Kitel, R., Pustelny, K., Gruba, N., Groborz, K., Jankowska, U., Kedracka-Krok, S., Wladyka, B., Drag, M., Lesner, A., Dubin, G.(2021) Int J Mol Sci 22
- PubMed: 33672341 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/ijms22042220
- Primary Citation Related Structures:
6SF7 - PubMed Abstract:
Accumulating evidence suggests that six proteases encoded in the spl operon of a dangerous human pathogen, Staphylococcus aureus , may play a role in virulence. Interestingly, SplA, B, D, and E have complementary substrate specificities while SplF remains to be characterized in this regard. Here, we describe the prerequisites of a heterologous expression system for active SplF protease and characterize the enzyme in terms of substrate specificity and its structural determinants. Substrate specificity of SplF is comprehensively profiled using combinatorial libraries of peptide substrates demonstrating strict preference for long aliphatic sidechains at the P1 subsite and significant selectivity for aromatic residues at P3. The crystal structure of SplF was provided at 1.7 Å resolution to define the structural basis of substrate specificity of SplF. The obtained results were compared and contrasted with the characteristics of other Spl proteases determined to date to conclude that the spl operon encodes a unique extracellular proteolytic system.
- Malopolska Center of Biotechnology, Jagiellonian University, 7a Gronostajowa St., 30-387 Krakow, Poland.
Organizational Affiliation: