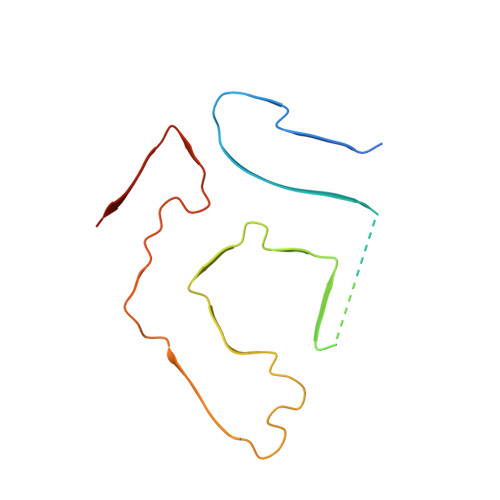
Cryo-EM structure of a transthyretin-derived amyloid fibril from a patient with hereditary ATTR amyloidosis.
Schmidt, M., Wiese, S., Adak, V., Engler, J., Agarwal, S., Fritz, G., Westermark, P., Zacharias, M., Fandrich, M.(2019) Nat Commun 10: 5008-5008
- PubMed: 31676763 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-019-13038-z
- Primary Citation Related Structures:
6SDZ - PubMed Abstract:
ATTR amyloidosis is one of the worldwide most abundant forms of systemic amyloidosis. The disease is caused by the misfolding of transthyretin protein and the formation of amyloid deposits at different sites within the body. Here, we present a 2.97 Å cryo electron microscopy structure of a fibril purified from the tissue of a patient with hereditary Val30Met ATTR amyloidosis. The fibril consists of a single protofilament that is formed from an N-terminal and a C-terminal fragment of transthyretin. Our structure provides insights into the mechanism of misfolding and implies the formation of an early fibril state from unfolded transthyretin molecules, which upon proteolysis converts into mature ATTR amyloid fibrils.
- Institute of Protein Biochemistry, Ulm University, 89081, Ulm, Germany.
Organizational Affiliation: