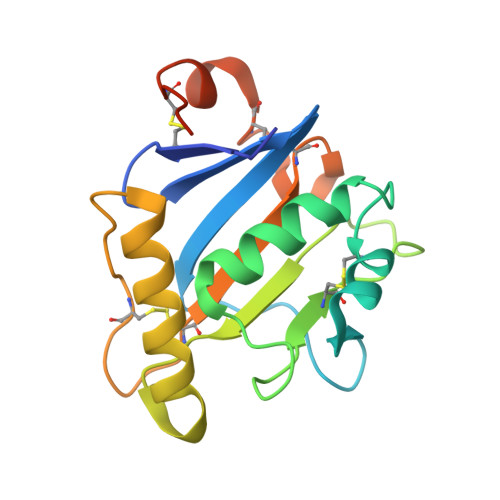
Rational design of universal immunotherapy for TfR1-tropic arenaviruses.
Cohen-Dvashi, H., Amon, R., Agans, K.N., Cross, R.W., Borenstein-Katz, A., Mateo, M., Baize, S., Padler-Karavani, V., Geisbert, T.W., Diskin, R.(2020) Nat Commun 11: 67-67
- PubMed: 31900422
- DOI: https://doi.org/10.1038/s41467-019-13924-6
- Primary Citation Related Structures:
6S9J - PubMed Abstract:
Certain arenaviruses that circulate in rodent populations can cause life-threatening hemorrhagic fevers when they infect humans. Due to their efficient transmission, arenaviruses pose a severe risk for outbreaks and might be exploited as biological weapons. Effective countermeasures against these viruses are highly desired. Ideally, a single remedy would be effective against many or even all the pathogenic viruses in this family. However, despite the fact that all pathogenic arenaviruses from South America utilize transferrin receptor 1 (TfR1) as a cellular receptor, their viral glycoproteins are highly diversified, impeding efforts to isolate cross-neutralizing antibodies. Here we address this problem using a rational design approach to target TfR1-tropic arenaviruses with high potency and breadth. The pan-reactive molecule is highly effective against all arenaviruses that were tested, offering a universal therapeutic approach. Our design scheme avoids the shortcomings of previous immunoadhesins and can be used to combat other zoonotic pathogens.
- Department of Structural Biology, Weizmann Institute of Science, Rehovot, 7610001, Israel.
Organizational Affiliation: