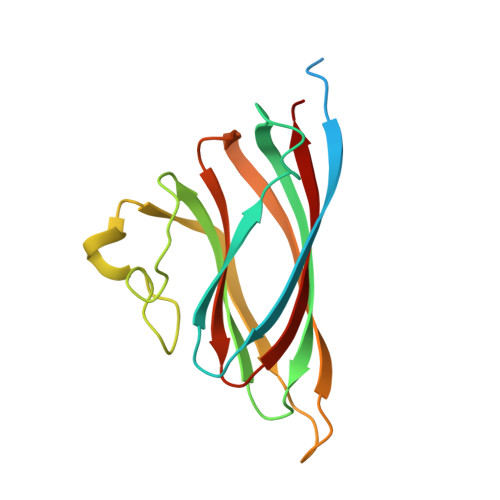
Structure-guided mutagenesis of the capsid protein indicates that a nanovirus requires assembled viral particles for systemic infection.
Trapani, S., Bhat, E.A., Yvon, M., Lai-Kee-Him, J., Hoh, F., Vernerey, M.S., Pirolles, E., Bonnamy, M., Schoehn, G., Zeddam, J.L., Blanc, S., Bron, P.(2023) PLoS Pathog 19: e1011086-e1011086
- PubMed: 36622854 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1011086
- Primary Citation Related Structures:
6S44 - PubMed Abstract:
Nanoviruses are plant multipartite viruses with a genome composed of six to eight circular single-stranded DNA segments. The distinct genome segments are encapsidated individually in icosahedral particles that measure ≈18 nm in diameter. Recent studies on the model species Faba bean necrotic stunt virus (FBNSV) revealed that complete sets of genomic segments rarely occur in infected plant cells and that the function encoded by a given viral segment can complement the others across neighbouring cells, presumably by translocation of the gene products through unknown molecular processes. This allows the viral genome to replicate, assemble into viral particles and infect anew, even with the distinct genome segments scattered in different cells. Here, we question the form under which the FBNSV genetic material propagates long distance within the vasculature of host plants and, in particular, whether viral particle assembly is required. Using structure-guided mutagenesis based on a 3.2 Å resolution cryogenic-electron-microscopy reconstruction of the FBNSV particles, we demonstrate that specific site-directed mutations preventing capsid formation systematically suppress FBNSV long-distance movement, and thus systemic infection of host plants, despite positive detection of the mutated coat protein when the corresponding segment is agroinfiltrated into plant leaves. These results strongly suggest that the viral genome does not propagate within the plant vascular system under the form of uncoated DNA molecules or DNA:coat-protein complexes, but rather moves long distance as assembled viral particles.
- CBS (Centre de Biologie Structurale), Univ Montpellier, CNRS, INSERM, Montpellier, France.
Organizational Affiliation: