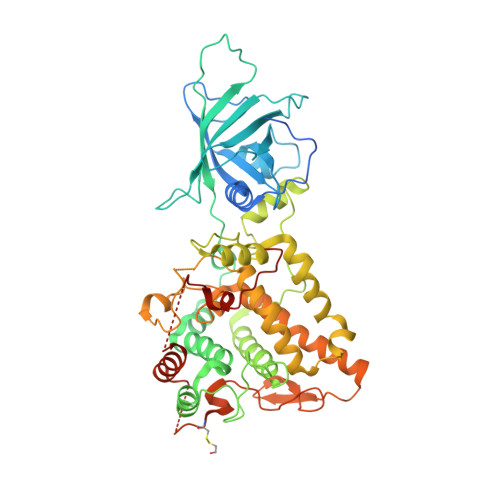
A novel mechanism of inhibition by phenylthiourea on PvdP, a tyrosinase synthesizing pyoverdine of Pseudomonas aeruginosa.
Wibowo, J.P., Batista, F.A., van Oosterwijk, N., Groves, M.R., Dekker, F.J., Quax, W.J.(2020) Int J Biol Macromol 146: 212-221
- PubMed: 31899238 Search on PubMed
- DOI: https://doi.org/10.1016/j.ijbiomac.2019.12.252
- Primary Citation Related Structures:
6RRP, 6RRQ, 6RRR - PubMed Abstract:
The biosynthesis of pyoverdine, the major siderophore of Pseudomonas aeruginosa, is a well-organized process involving a discrete number of enzyme-catalyzed steps. The final step of this process involves the PvdP tyrosinase, which converts ferribactin into pyoverdine. Thus, inhibition of the PvdP tyrosinase activity provides an attractive strategy to interfere with siderophore synthesis to manage P. aeruginosa infections. Here, we report phenylthiourea as a non-competitive inhibitor of PvdP for which we solved a crystal structure in complex with PvdP. The crystal structure indicates that phenylthiourea binds to an allosteric binding site and thereby interferes with its tyrosinase activity. We further provide proofs that PvdP tyrosinase inhibition by phenylthiourea requires the C-terminal lid region. This provides opportunities to develop inhibitors that target the allosteric site, which seems to be confined to fluorescent pseudomonads, and not the tyrosinase active site. Furthermore, increases the chances to identify PvdP inhibitors that selectively interfere with siderophore synthesis.
- Department of Chemical and Pharmaceutical Biology, Groningen Research Institute of Pharmacy, University of Groningen, the Netherlands; Faculty of Pharmacy, University of Muhammadiyah Banjarmasin, Banjarmasin, Indonesia.
Organizational Affiliation: