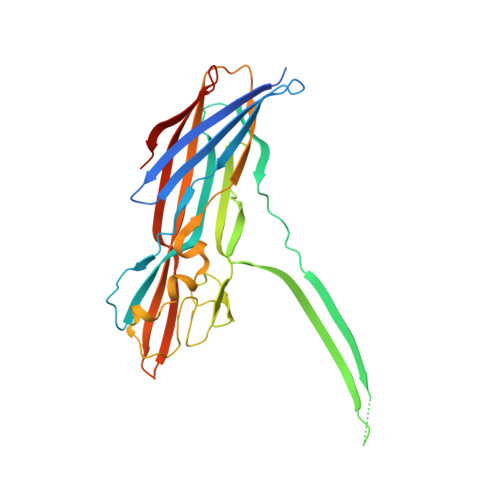
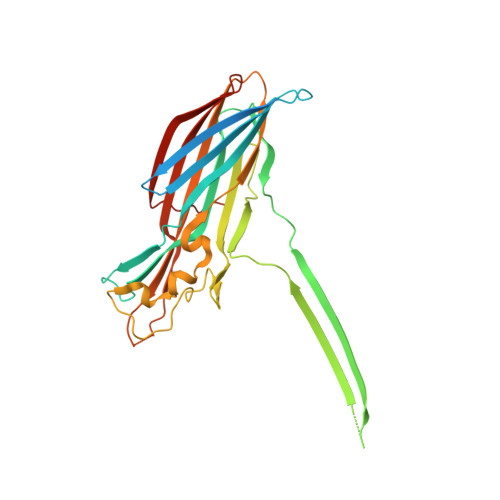
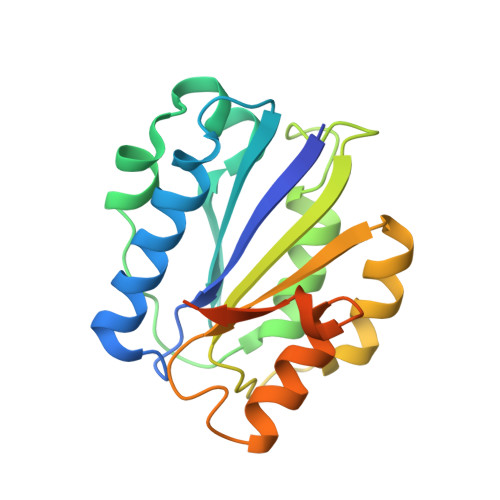
Molecular mechanism of leukocidin GH-integrin CD11b/CD18 recognition and species specificity.
Trstenjak, N., Milic, D., Graewert, M.A., Rouha, H., Svergun, D., Djinovic-Carugo, K., Nagy, E., Badarau, A.(2020) Proc Natl Acad Sci U S A 117: 317-327
- PubMed: 31852826 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1913690116
- Primary Citation Related Structures:
6RHV, 6RHW - PubMed Abstract:
Host-pathogen interactions are central to understanding microbial pathogenesis. The staphylococcal pore-forming cytotoxins hijack important immune molecules but little is known about the underlying molecular mechanisms of cytotoxin-receptor interaction and host specificity. Here we report the structures of a staphylococcal pore-forming cytotoxin, leukocidin GH (LukGH), in complex with its receptor (the α-I domain of complement receptor 3, CD11b-I), both for the human and murine homologs. We observe 2 binding interfaces, on the LukG and the LukH protomers, and show that human CD11b-I induces LukGH oligomerization in solution. LukGH binds murine CD11b-I weakly and is inactive toward murine neutrophils. Using a LukGH variant engineered to bind mouse CD11b-I, we demonstrate that cytolytic activity does not only require binding but also receptor-dependent oligomerization. Our studies provide an unprecedented insight into bicomponent leukocidin-host receptor interaction, enabling the development of antitoxin approaches and improved animal models to explore these approaches.
- Arsanis Biosciences, Vienna Biocenter, 1030 Vienna, Austria.
Organizational Affiliation: