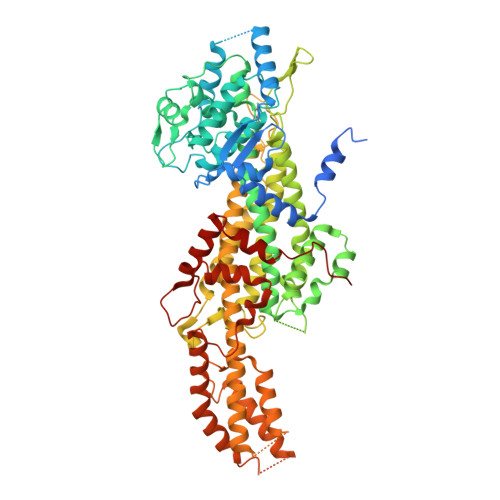
Mapping the Hydrophobic Substrate Binding Site of Phenylalanine Ammonia Lyase from Petroselinum crispum
Nagy, E.Z.A., Tork, S.D., Lang, P.A., Filip, A., Irimie, F.D., Poppe, L., Tosa, M.I., Schofield, C.J., Brem, J., Paizs, C., Bencze, L.C.(2019) ACS Catal