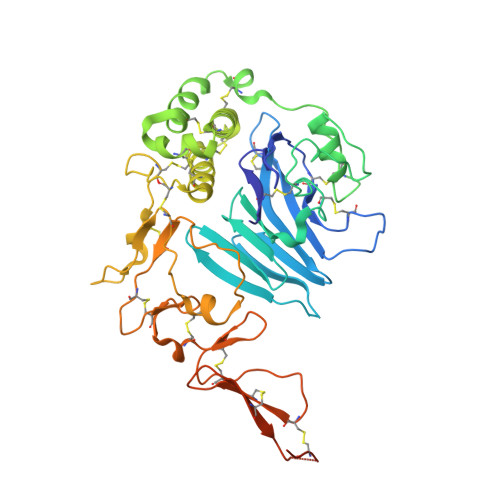
Intestinal Gel-Forming Mucins Polymerize by Disulfide-Mediated Dimerization of D3 Domains.
Javitt, G., Calvo, M.L.G., Albert, L., Reznik, N., Ilani, T., Diskin, R., Fass, D.(2019) J Mol Biology 431: 3740-3752
- PubMed: 31310764 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2019.07.018
- Primary Citation Related Structures:
6RBF - PubMed Abstract:
The mucin 2 glycoprotein assembles into a complex hydrogel that protects intestinal epithelia and houses the gut microbiome. A major step in mucin 2 assembly is further multimerization of preformed mucin dimers, thought to produce a honeycomb-like arrangement upon hydrogel expansion. Important open questions are how multiple mucin 2 dimers become covalently linked to one another and how mucin 2 multimerization compares with analogous processes in related polymers such as respiratory tract mucins and the hemostasis protein von Willebrand factor. Here we report the x-ray crystal structure of the mucin 2 multimerization module, found to form a dimer linked by two intersubunit disulfide bonds. The dimer structure calls into question the current model for intestinal mucin assembly, which proposes disulfide-mediated trimerization of the same module. Key residues making interactions across the dimer interface are highly conserved in intestinal mucin orthologs, supporting the physiological relevance of the observed quaternary structure. With knowledge of the interface residues, it can be demonstrated that many of these amino acids are also present in other mucins and in von Willebrand factor, further indicating that the stable dimer arrangement reported herein is likely to be shared across this functionally broad protein family. The mucin 2 module structure thus reveals the manner by which both mucins and von Willebrand factor polymerize, drawing deep structural parallels between macromolecular assemblies critical to mucosal epithelia and the vasculature.
- Department of Structural Biology, Weizmann Institute of Science, Rehovot 7610001, Israel.
Organizational Affiliation: