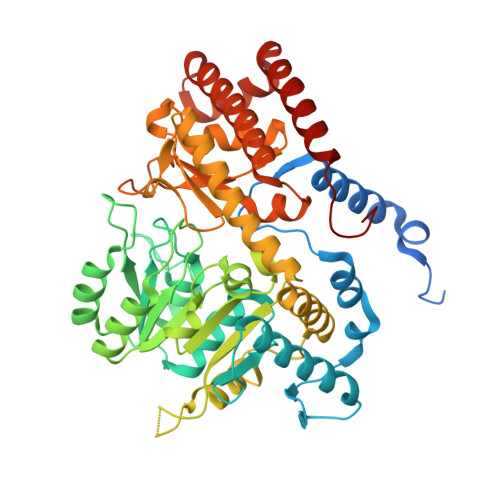
Structural basis of inhibition of the human serine hydroxymethyltransferase SHMT2 by antifolate drugs.
Scaletti, E., Jemth, A.S., Helleday, T., Stenmark, P.(2019) FEBS Lett 593: 1863-1873
- PubMed: 31127856 Search on PubMed
- DOI: https://doi.org/10.1002/1873-3468.13455
- Primary Citation Related Structures:
6QVG, 6QVL - PubMed Abstract:
Serine hydroxymethyltransferase (SHMT) is the major source of 1-carbon units required for nucleotide synthesis. Humans have cytosolic (SHMT1) and mitochondrial (SHMT2) isoforms, which are upregulated in numerous cancers, making the enzyme an attractive drug target. Here, we show that the antifolates lometrexol and pemetrexed are inhibitors of SHMT2 and solve the first SHMT2-antifolate structures. The antifolates display large differences in their hydrogen bond networks despite their similarity. Lometrexol was found to be the best hSHMT1/2 inhibitor from a panel antifolates. Comparison of apo hSHMT1 with antifolate bound hSHMT2 indicates a highly conserved active site architecture. This structural information offers insights as to how these compounds could be improved to produce more potent and specific inhibitors of this emerging anti-cancer drug target.
- Department of Biochemistry and Biophysics, Stockholm University, Sweden.
Organizational Affiliation: