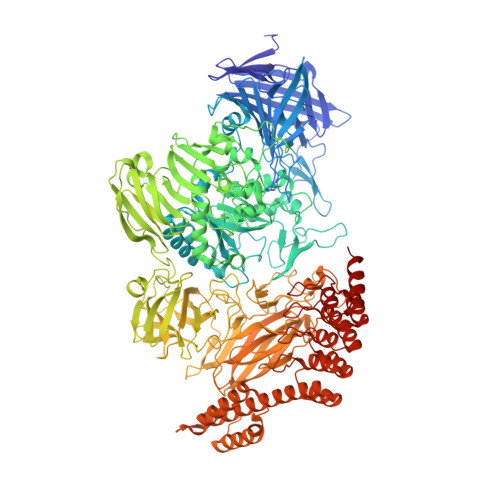
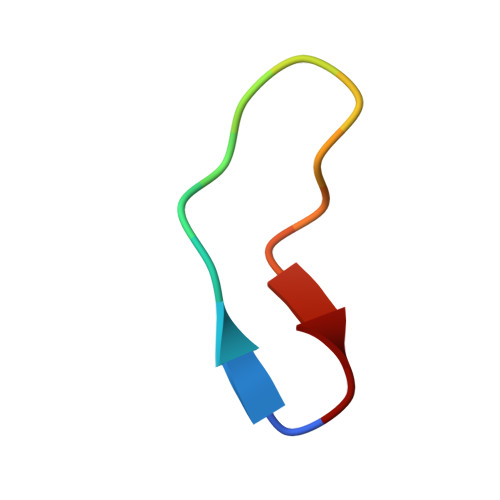
Structural analysis of biological targets by host:guest crystal lattice engineering.
Ernst, P., Pluckthun, A., Mittl, P.R.E.(2019) Sci Rep 9: 15199-15199
- PubMed: 31645583 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-019-51017-y
- Primary Citation Related Structures:
6QEP, 6QEV, 6QFK, 6QFO, 6SH9 - PubMed Abstract:
To overcome the laborious identification of crystallisation conditions for protein X-ray crystallography, we developed a method where the examined protein is immobilised as a guest molecule in a universal host lattice. We applied crystal engineering to create a generic crystalline host lattice under reproducible, predefined conditions and analysed the structures of target guest molecules of different size, namely two 15-mer peptides and green fluorescent protein (sfGFP). A fusion protein with an N-terminal endo-α-N-acetylgalactosaminidase (EngBF) domain and a C-terminal designed ankyrin repeat protein (DARPin) domain establishes the crystal lattice. The target is recruited into the host lattice, always in the same crystal form, through binding to the DARPin. The target structures can be determined rapidly from difference Fourier maps, whose quality depends on the size of the target and the orientation of the DARPin.
- Department of Biochemistry, University of Zürich, Winterthurerstrasse 190, CH-8057, Zürich, Switzerland.
Organizational Affiliation: