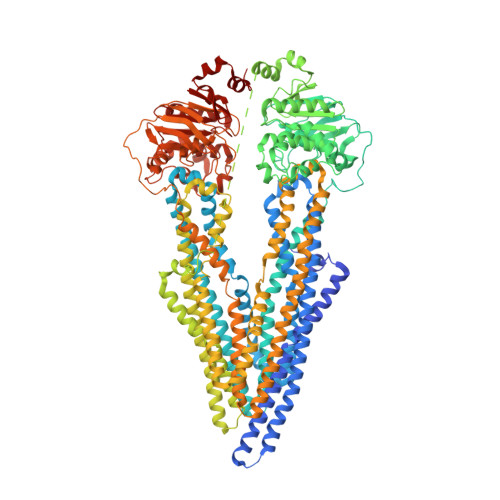
Structural insight into substrate and inhibitor discrimination by human P-glycoprotein.
Alam, A., Kowal, J., Broude, E., Roninson, I., Locher, K.P.(2019) Science 363: 753-756
- PubMed: 30765569 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.aav7102
- Primary Citation Related Structures:
6QEE, 6QEX - PubMed Abstract:
ABCB1, also known as P-glycoprotein, actively extrudes xenobiotic compounds across the plasma membrane of diverse cells, which contributes to cellular drug resistance and interferes with therapeutic drug delivery. We determined the 3.5-angstrom cryo-electron microscopy structure of substrate-bound human ABCB1 reconstituted in lipidic nanodiscs, revealing a single molecule of the chemotherapeutic compound paclitaxel (Taxol) bound in a central, occluded pocket. A second structure of inhibited, human-mouse chimeric ABCB1 revealed two molecules of zosuquidar occupying the same drug-binding pocket. Minor structural differences between substrate- and inhibitor-bound ABCB1 sites are amplified toward the nucleotide-binding domains (NBDs), revealing how the plasticity of the drug-binding site controls the dynamics of the adenosine triphosphate-hydrolyzing NBDs. Ordered cholesterol and phospholipid molecules suggest how the membrane modulates the conformational changes associated with drug binding and transport.
- Institute of Molecular Biology and Biophysics, ETH Zürich, Otto-Stern-Weg 5, 8093 Zürich, Switzerland.
Organizational Affiliation: