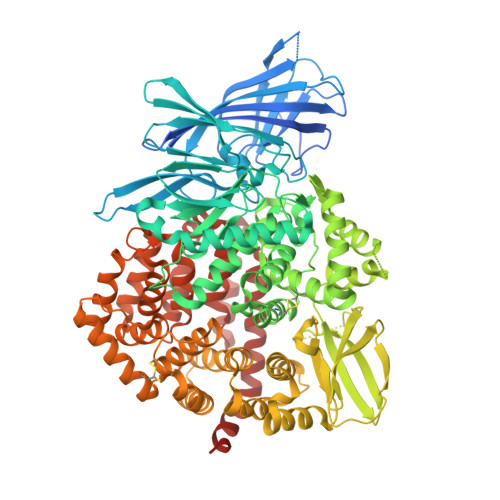
High-Resolution Crystal Structure of Endoplasmic Reticulum Aminopeptidase 1 with Bound Phosphinic Transition-State Analogue Inhibitor.
Giastas, P., Neu, M., Rowland, P., Stratikos, E.(2019) ACS Med Chem Lett 10: 708-713
- PubMed: 31097987 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsmedchemlett.9b00002
- Primary Citation Related Structures:
6Q4R - PubMed Abstract:
Endoplasmic reticulum aminopeptidase 1 (ERAP1) is an intracellular enzyme that helps generate peptides presented by Major Histocompatibility Complex Class I (MHC class I) molecules and is an emerging target for immunotherapy applications. Despite almost two decades of research on ERAP1, lack of high-resolution crystal structures has hampered drug-development efforts. By optimizing the protein construct, we obtained a high-resolution (1.60 Å) crystal structure of the closed-conformation of ERAP1 with a potent phosphinic pseudopeptide inhibitor bound in its active site. The structure provides key insight on the mechanism of inhibition as well as selectivity toward homologous enzymes and allows detailed mapping of the internal cavity of the enzyme that accommodates peptide-substrates. Bis-tris propane and malic acid molecules, found bound in pockets in the internal cavity, reveal potential druggable secondary binding sites. The ability to obtain high-resolution crystal structures of ERAP1 removes a major bottleneck in the development of compounds that regulate its activity and will greatly accelerate drug-discovery efforts.
- National Center for Scientific Research Demokritos, Agia Paraskevi, Athens 15310, Greece.
Organizational Affiliation: