Mutagenesis and Structural Studies Reveal the Basis for the Activity and Stability Properties That Distinguish thePhotinusLuciferasesscintillansandpyralis.
Branchini, B.R., Fontaine, D.M., Southworth, T.L., Huta, B.P., Racela, A., Patel, K.D., Gulick, A.M.(2019) Biochemistry 58: 4293-4303
- PubMed: 31560532 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.9b00719
- Primary Citation Related Structures:
6Q2M - PubMed Abstract:
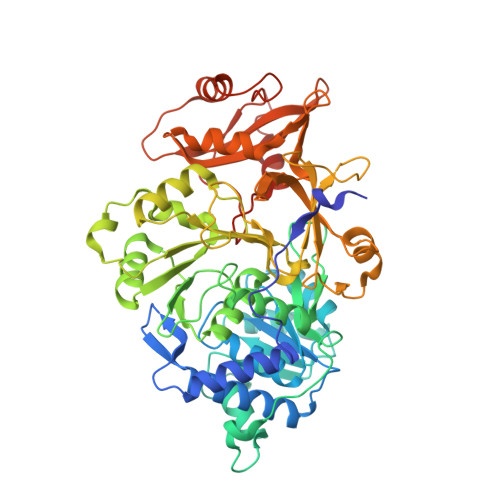
The dazzling yellow-green light emission of the common North American firefly Photinus pyralis and other bioluminescent organisms has provided a wide variety of prominent research applications like reporter gene assays and in vivo imaging methods. While the P. pyralis enzyme has been extensively studied, only recently has a second Photinus luciferase been cloned from the species scintillans . Even though the enzymes share very high sequence identity (89.8%), the color of the light they emit, their specific activity and their stability to heat, pH, and chemical denaturation are quite different with the scintillans luciferase being generally more resistant. Through the construction and evaluation of the properties of chimeric domain swapped, single point, and various combined variants, we have determined that only six amino acid changes are necessary to confer all of the properties of the scintillans enzyme to wild-type P. pyralis luciferase. Altered stability properties were attributed to four of the amino acid changes (T214N/S276T/H332N/E354N), and single mutations each predominantly changed emission color (Y255F) and specific activity (A222C). Results of a crystallographic study of the P. pyralis enzyme containing the six changes (Pps6) provide some insight into the structural basis for some of the documented property differences.
- Department of Chemistry , Connecticut College , New London , Connecticut 06320 , United States.
Organizational Affiliation: