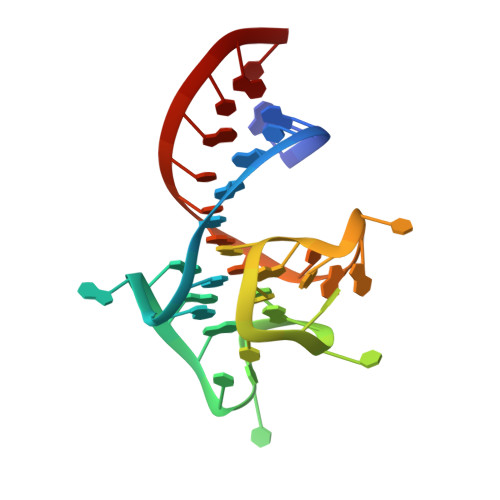
Co-crystal structure of the iMango-III fluorescent RNA aptamer using an X-ray free-electron laser.
Trachman III, R.J., Stagno, J.R., Conrad, C., Jones, C.P., Fischer, P., Meents, A., Wang, Y.X., Ferre-D'Amare, A.R.(2019) Acta Crystallogr F Struct Biol Commun 75: 547-551
- PubMed: 31397326 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X19010136
- Primary Citation Related Structures:
6PQ7 - PubMed Abstract:
Turn-on aptamers are in vitro-selected RNAs that bind to conditionally fluorescent small molecules and enhance their fluorescence. Upon binding TO1-biotin, the iMango-III aptamer achieves the largest fluorescence enhancement reported for turn-on aptamers (over 5000-fold). This aptamer was generated by structure-guided engineering and functional reselection of the parental aptamer Mango-III. Structures of both Mango-III and iMango-III have previously been determined by conventional cryocrystallography using synchrotron X-radiation. Using an X-ray free-electron laser (XFEL), the room-temperature iMango-III-TO1-biotin co-crystal structure has now been determined at 3.0 Å resolution. This structural model, which was refined against a data set of ∼1300 diffraction images (each from a single crystal), is largely consistent with the structures determined from single-crystal data sets collected at 100 K. This constitutes a technical benchmark on the way to XFEL pump-probe experiments on fluorescent RNA-small molecule complexes.
- Biochemistry and Biophysics Center, National Heart, Lung and Blood Institute, Bethesda, Maryland, USA.
Organizational Affiliation: